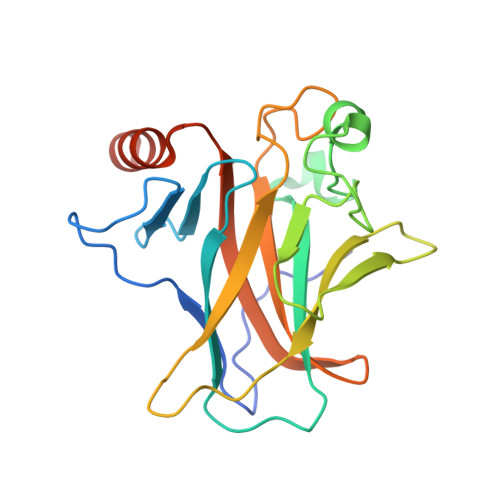
Evolutionary history of the p53 family DNA-binding domain: insights from an Alvinella pompejana homolog.
Zhang, Q., Balourdas, D.I., Baron, B., Senitzki, A., Haran, T.E., Wiman, K.G., Soussi, T., Joerger, A.C.(2022) Cell Death Dis 13: 214-214
- PubMed: 35256607
- DOI: https://doi.org/10.1038/s41419-022-04653-8
- Primary Citation of Related Structures:
7PC6 - PubMed Abstract:
The extremophile Alvinella pompejana, an annelid worm living on the edge of hydrothermal vents in the Pacific Ocean, is an excellent model system for studying factors that govern protein stability. Low intrinsic stability is a crucial factor for the susceptibility of the transcription factor p53 to inactivating mutations in human cancer. Understanding its molecular basis may facilitate the design of novel therapeutic strategies targeting mutant p53. By analyzing expressed sequence tag (EST) data, we discovered a p53 family gene in A. pompejana. Protein crystallography and biophysical studies showed that it has a p53/p63-like DNA-binding domain (DBD) that is more thermostable than all vertebrate p53 DBDs tested so far, but not as stable as that of human p63. We also identified features associated with its increased thermostability. In addition, the A. pompejana homolog shares DNA-binding properties with human p53 family DBDs, despite its evolutionary distance, consistent with a potential role in maintaining genome integrity. Through extensive structural and phylogenetic analyses, we could further trace key evolutionary events that shaped the structure, stability, and function of the p53 family DBD over time, leading to a potent but vulnerable tumor suppressor in humans.
Organizational Affiliation:
Department of Neuroscience, Biomedicum, Karolinska Institutet, Stockholm, Sweden.