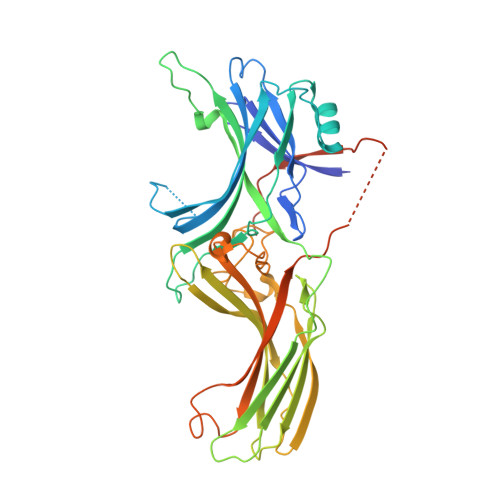
Structural evidence for visual arrestin priming via complexation of phosphoinositols.
Sander, C.L., Luu, J., Kim, K., Furkert, D., Jang, K., Reichenwallner, J., Kang, M., Lee, H.J., Eger, B.T., Choe, H.W., Fiedler, D., Ernst, O.P., Kim, Y.J., Palczewski, K., Kiser, P.D.(2022) Structure 30: 263-277.e5
- PubMed: 34678158
- DOI: https://doi.org/10.1016/j.str.2021.10.002
- Primary Citation of Related Structures:
7F1W, 7F1X, 7JSM, 7JTB, 7JXA, 7MOR, 7MP0, 7MP1, 7MP2 - PubMed Abstract:
Visual arrestin (Arr1) terminates rhodopsin signaling by blocking its interaction with transducin. To do this, Arr1 translocates from the inner to the outer segment of photoreceptors upon light stimulation. Mounting evidence indicates that inositol phosphates (InsPs) affect Arr1 activity, but the Arr1-InsP molecular interaction remains poorly defined. We report the structure of bovine Arr1 in a ligand-free state featuring a near-complete model of the previously unresolved C-tail, which plays a crucial role in regulating Arr1 activity. InsPs bind to the N-domain basic patch thus displacing the C-tail, suggesting that they prime Arr1 for interaction with rhodopsin and help direct Arr1 translocation. These structures exhibit intact polar cores, suggesting that C-tail removal by InsP binding is insufficient to activate Arr1. These results show how Arr1 activity can be controlled by endogenous InsPs in molecular detail.
Organizational Affiliation:
Department of Pharmacology, Case Western Reserve University, Cleveland, OH 44106, USA; Department of Ophthalmology and the Gavin Herbert Eye Institute, University of California, Irvine, CA 92697, USA.