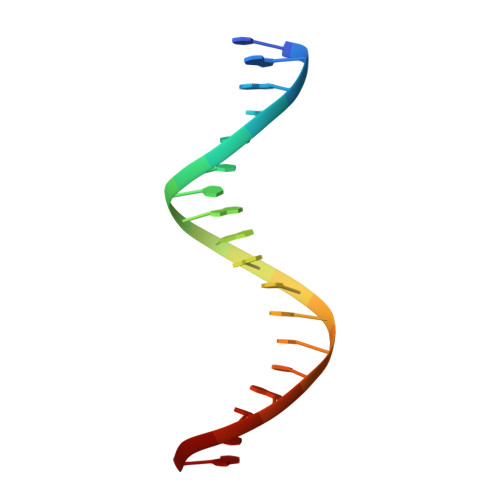
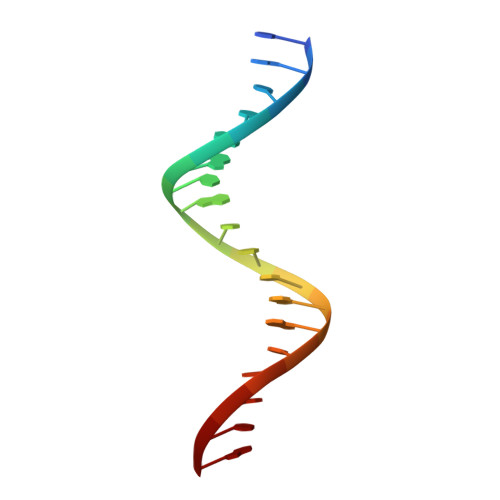
Structural basis of direct and inverted DNA sequence repeat recognition by helix-turn-helix transcription factors.
Fernandez-Lopez, R., Ruiz, R., Del Campo, I., Gonzalez-Montes, L., Boer, D.R., de la Cruz, F., Moncalian, G.(2022) Nucleic Acids Res 50: 11938-11947
- PubMed: 36370103
- DOI: https://doi.org/10.1093/nar/gkac1024
- Primary Citation of Related Structures:
7BBQ, 7BCA, 7BCB - PubMed Abstract:
Some transcription factors bind DNA motifs containing direct or inverted sequence repeats. Preference for each of these DNA topologies is dictated by structural constraints. Most prokaryotic regulators form symmetric oligomers, which require operators with a dyad structure. Binding to direct repeats requires breaking the internal symmetry, a property restricted to a few regulators, most of them from the AraC family. The KorA family of transcriptional repressors, involved in plasmid propagation and stability, includes members that form symmetric dimers and recognize inverted repeats. Our structural analyses show that ArdK, a member of this family, can form a symmetric dimer similar to that observed for KorA, yet it binds direct sequence repeats as a non-symmetric dimer. This is possible by the 180° rotation of one of the helix-turn-helix domains. We then probed and confirmed that ArdK shows affinity for an inverted repeat, which, surprisingly, is also recognized by a non-symmetrical dimer. Our results indicate that structural flexibility at different positions in the dimerization interface constrains transcription factors to bind DNA sequences with one of these two alternative DNA topologies.
Organizational Affiliation:
Departamento de Biología Molecular, Universidad de Cantabria and Instituto de Biomedicina y Biotecnología de Cantabria (IBBTEC), CSIC-Universidad de Cantabria, 39011, Santander, Spain.