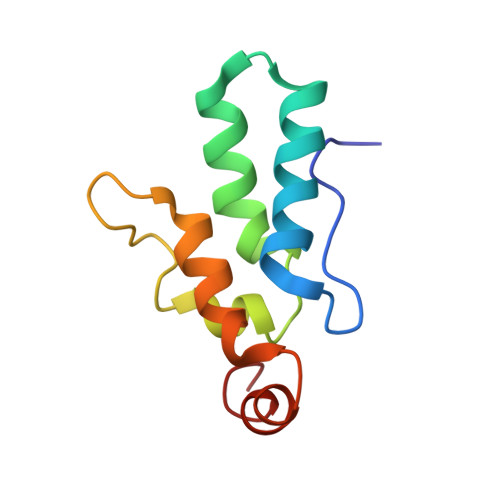
Structural basis for Fullerene geometry in a human endogenous retrovirus capsid.
Acton, O., Grant, T., Nicastro, G., Ball, N.J., Goldstone, D.C., Robertson, L.E., Sader, K., Nans, A., Ramos, A., Stoye, J.P., Taylor, I.A., Rosenthal, P.B.(2019) Nat Commun 10: 5822-5822
- PubMed: 31862888
- DOI: https://doi.org/10.1038/s41467-019-13786-y
- Primary Citation of Related Structures:
6SA9, 6SAI, 6SSJ, 6SSK, 6SSL, 6SSM - PubMed Abstract:
The HML2 (HERV-K) group constitutes the most recently acquired family of human endogenous retroviruses, with many proviruses less than one million years old. Many maintain intact open reading frames and provirus expression together with HML2 particle formation are observed in early stage human embryo development and are associated with pluripotency as well as inflammatory disease, cancers and HIV-1 infection. Here, we reconstruct the core structural protein (CA) of an HML2 retrovirus, assemble particles in vitro and employ single particle cryogenic electron microscopy (cryo-EM) to determine structures of four classes of CA Fullerene shell assemblies. These icosahedral and capsular assemblies reveal at high-resolution the molecular interactions that allow CA to form both pentamers and hexamers and show how invariant pentamers and structurally plastic hexamers associate to form the unique polyhedral structures found in retroviral cores.
Organizational Affiliation:
Structural Biology of Cells and Viruses Laboratory, The Francis Crick Institute, 1 Midland Road, London, NW1 1AT, UK.