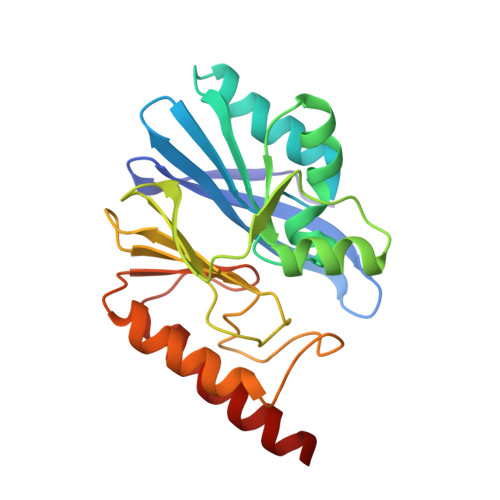
Structure and Molecular Recognition Mechanism of IMP-13 Metallo-beta-Lactamase.
Softley, C.A., Zak, K.M., Bostock, M.J., Fino, R., Zhou, R.X., Kolonko, M., Mejdi-Nitiu, R., Meyer, H., Sattler, M., Popowicz, G.M.(2020) Antimicrob Agents Chemother 64
- PubMed: 32205343
- DOI: https://doi.org/10.1128/AAC.00123-20
- Primary Citation of Related Structures:
6R73, 6R78, 6R79, 6RZR, 6RZS, 6S0H - PubMed Abstract:
Multidrug resistance among Gram-negative bacteria is a major global public health threat. Metallo-β-lactamases (MBLs) target the most widely used antibiotic class, the β-lactams, including the most recent generation of carbapenems. Interspecies spread renders these enzymes a serious clinical threat, and there are no clinically available inhibitors. We present the crystal structures of IMP-13, a structurally uncharacterized MBL from the Gram-negative bacterium Pseudomonas aeruginosa found in clinical outbreaks globally, and characterize the binding using solution nuclear magnetic resonance spectroscopy and molecular dynamics simulations. The crystal structures of apo IMP-13 and IMP-13 bound to four clinically relevant carbapenem antibiotics (doripenem, ertapenem, imipenem, and meropenem) are presented. Active-site plasticity and the active-site loop, where a tryptophan residue stabilizes the antibiotic core scaffold, are essential to the substrate-binding mechanism. The conserved carbapenem scaffold plays the most significant role in IMP-13 binding, explaining the broad substrate specificity. The observed plasticity and substrate-locking mechanism provide opportunities for rational drug design of novel metallo-β-lactamase inhibitors, essential in the fight against antibiotic resistance.
Organizational Affiliation:
Biomolecular NMR and Center for Integrated Protein Science Munich at Department Chemie, Technical University of Munich, Garching, Germany.