Structural basis for Marburg virus neutralization by a cross-reactive human antibody.
Hashiguchi, T., Fusco, M.L., Bornholdt, Z.A., Lee, J.E., Flyak, A.I., Matsuoka, R., Kohda, D., Yanagi, Y., Hammel, M., Crowe, J.E., Saphire, E.O.(2015) Cell 160: 904-912
- PubMed: 25723165
- DOI: https://doi.org/10.1016/j.cell.2015.01.041
- Primary Citation of Related Structures:
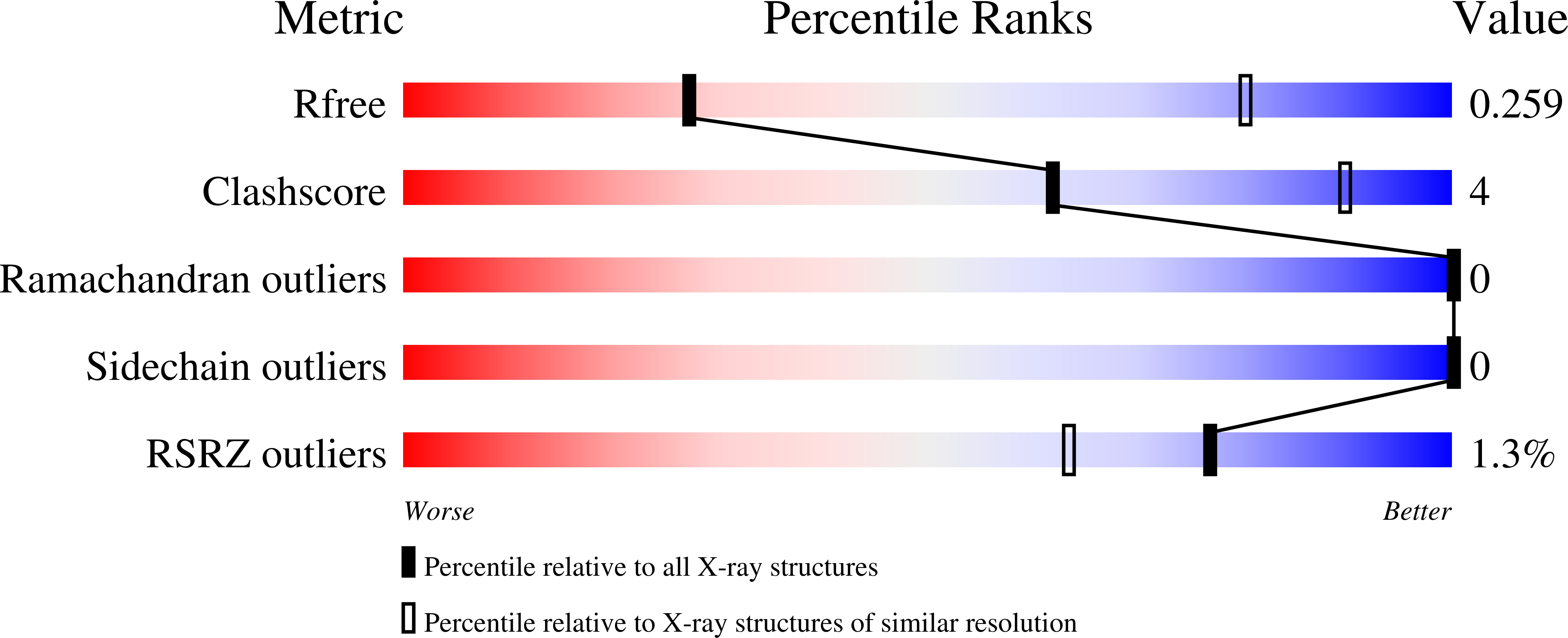
5UQY - PubMed Abstract:
The filoviruses, including Marburg and Ebola, express a single glycoprotein on their surface, termed GP, which is responsible for attachment and entry of target cells. Filovirus GPs differ by up to 70% in protein sequence, and no antibodies are yet described that cross-react among them. Here, we present the 3.6 Å crystal structure of Marburg virus GP in complex with a cross-reactive antibody from a human survivor, and a lower resolution structure of the antibody bound to Ebola virus GP. The antibody, MR78, recognizes a GP1 epitope conserved across the filovirus family, which likely represents the binding site of their NPC1 receptor. Indeed, MR78 blocks binding of the essential NPC1 domain C. These structures and additional small-angle X-ray scattering of mucin-containing MARV and EBOV GPs suggest why such antibodies were not previously elicited in studies of Ebola virus, and provide critical templates for development of immunotherapeutics and inhibitors of entry.
Organizational Affiliation:
Department of Immunology and Microbial Science, The Scripps Research Institute, La Jolla, CA 92037, USA; Department of Virology, Faculty of Medicine, Kyushu University, Fukuoka, 812-8582, Japan.