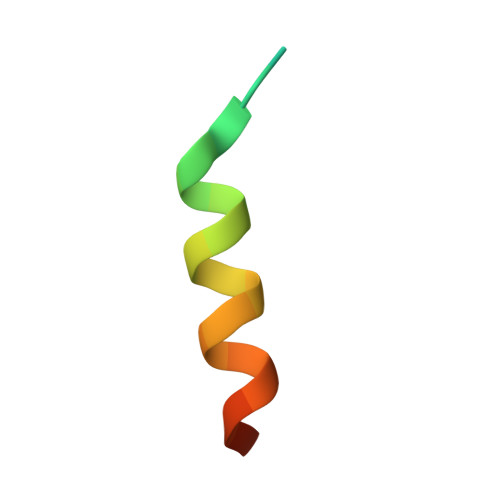
Distinct Mechanisms of Recognizing Endosomal Sorting Complex Required for Transport III (ESCRT-III) Protein IST1 by Different Microtubule Interacting and Trafficking (MIT) Domains.
Guo, E.Z., Xu, Z.(2015) J Biol Chem 290: 8396-8408
- PubMed: 25657007
- DOI: https://doi.org/10.1074/jbc.M114.607903
- Primary Citation of Related Structures:
4U7E, 4U7I, 4U7Y - PubMed Abstract:
The endosomal sorting complex required for transport (ESCRT) machinery is responsible for membrane remodeling in a number of biological processes including multivesicular body biogenesis, cytokinesis, and enveloped virus budding. In mammalian cells, efficient abscission during cytokinesis requires proper function of the ESCRT-III protein IST1, which binds to the microtubule interacting and trafficking (MIT) domains of VPS4, LIP5, and Spartin via its C-terminal MIT-interacting motif (MIM). Here, we studied the molecular interactions between IST1 and the three MIT domain-containing proteins to understand the structural basis that governs pairwise MIT-MIM interaction. Crystal structures of the three molecular complexes revealed that IST1 binds to the MIT domains of VPS4, LIP5, and Spartin using two different mechanisms (MIM1 mode versus MIM3 mode). Structural comparison revealed that structural features in both MIT and MIM contribute to determine the specific binding mechanism. Within the IST1 MIM sequence, two phenylalanine residues were shown to be important in discriminating MIM1 versus MIM3 binding. These observations enabled us to deduce a preliminary binding code, which we applied to provide CHMP2A, a protein that normally only binds the MIT domain in the MIM1 mode, the additional ability to bind the MIT domain of Spartin in the MIM3 mode.
Organizational Affiliation:
From the Life Science Institute and.