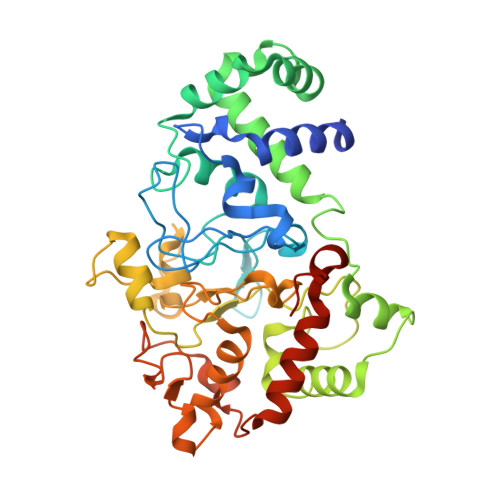
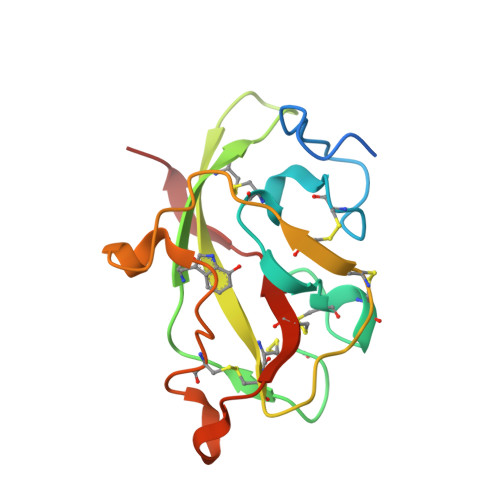
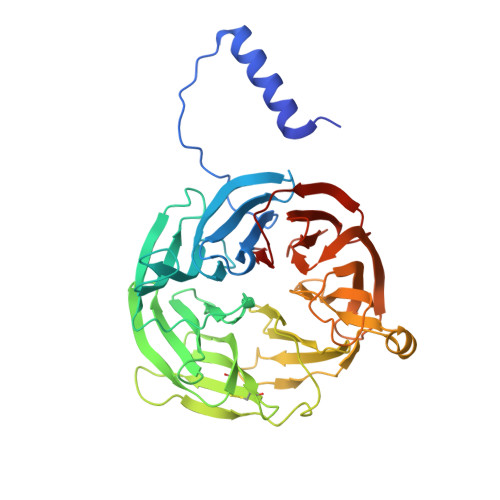
Site-directed mutagenesis of Gln103 reveals the influence of this residue on the redox properties and stability of MauG.
Shin, S., Yukl, E.T., Sehanobish, E., Wilmot, C.M., Davidson, V.L.(2014) Biochemistry 53: 1342-1349
- PubMed: 24517455
- DOI: https://doi.org/10.1021/bi5000349
- Primary Citation of Related Structures:
4O1Q - PubMed Abstract:
The diheme enzyme MauG catalyzes a six-electron oxidation that is required for the posttranslational modification of a precursor of methylamine dehydrogenase (preMADH) to complete the biosynthesis of its protein-derived cofactor, tryptophan tryptophylquinone (TTQ). Crystallographic and computational studies have implicated Gln103 in stabilizing the Fe(IV)═O moiety of the bis-Fe(IV) state by hydrogen bonding. The role of Gln103 was probed by site-directed mutagenesis. Q103L and Q103E mutations resulted in no expression and very little expression of the protein, respectively. Q103A MauG exhibited oxidative damage when isolated. Q103N MauG was isolated at levels comparable to that of wild-type MauG and exhibited normal activity in catalyzing the biosynthesis of TTQ from preMADH. The crystal structure of the Q103N MauG-preMADH complex suggests that a water may mediate hydrogen bonding between the shorter Asn103 side chain and the Fe(IV)═O moiety. The Q103N mutation caused the two redox potentials associated with the diferric/diferrous redox couple to become less negative, although the redox cooperativity of the hemes of MauG was retained. Upon addition of H2O2, Q103N MauG exhibits changes in the absorbance spectrum in the Soret and near-IR regions consistent with formation of the bis-Fe(IV) redox state. However, the rate of spontaneous return of the spectrum in the Soret region was 4.5-fold greater for Q103N MauG than for wild-type MauG. In contrast, the rate of spontaneous decay of the absorbance at 950 nm, which is associated with charge-resonance stabilization of the high-valence state, was similar for wild-type MauG and Q103N MauG. This suggests that as a consequence of the mutation a different distribution of resonance structures stabilizes the bis-Fe(IV) state. These results demonstrate that subtle changes in the structure of the side chain of residue 103 can significantly affect the overall protein stability of MauG and alter the redox properties of the hemes.
Organizational Affiliation:
Burnett School of Biomedical Sciences, College of Medicine, University of Central Florida , Orlando, Florida 32827, United States.