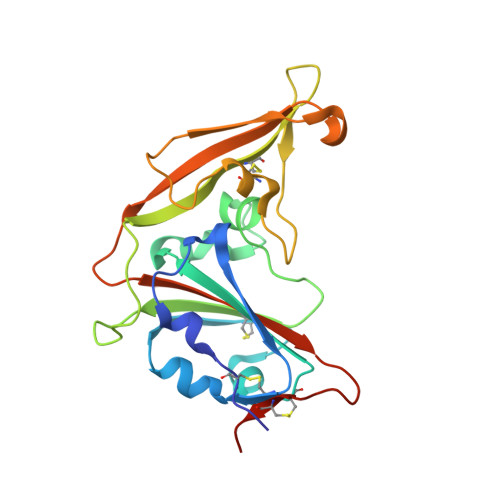
Crystal structure of the receptor-binding domain from newly emerged middle East respiratory syndrome coronavirus.
Chen, Y., Rajashankar, K.R., Yang, Y., Agnihothram, S.S., Liu, C., Lin, Y.L., Baric, R.S., Li, F.(2013) J Virol 87: 10777-10783
- PubMed: 23903833
- DOI: https://doi.org/10.1128/JVI.01756-13
- Primary Citation of Related Structures:
4L3N - PubMed Abstract:
The newly emerged Middle East respiratory syndrome coronavirus (MERS-CoV) has infected at least 77 people, with a fatality rate of more than 50%. Alarmingly, the virus demonstrates the capability of human-to-human transmission, raising the possibility of global spread and endangering world health and economy. Here we have identified the receptor-binding domain (RBD) from the MERS-CoV spike protein and determined its crystal structure. This study also presents a structural comparison of MERS-CoV RBD with other coronavirus RBDs, successfully positioning MERS-CoV on the landscape of coronavirus evolution and providing insights into receptor binding by MERS-CoV. Furthermore, we found that MERS-CoV RBD functions as an effective entry inhibitor of MERS-CoV. The identified MERS-CoV RBD may also serve as a potential candidate for MERS-CoV subunit vaccines. Overall, this study enhances our understanding of the evolution of coronavirus RBDs, provides insights into receptor recognition by MERS-CoV, and may help control the transmission of MERS-CoV in humans.
Organizational Affiliation:
Department of Pharmacology, University of Minnesota Medical School, Minneapolis, Minnesota, USA.