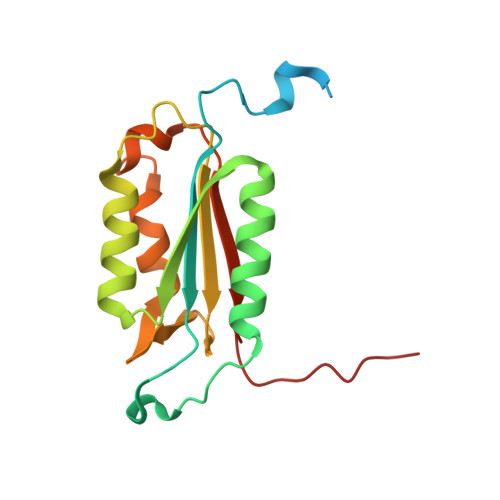
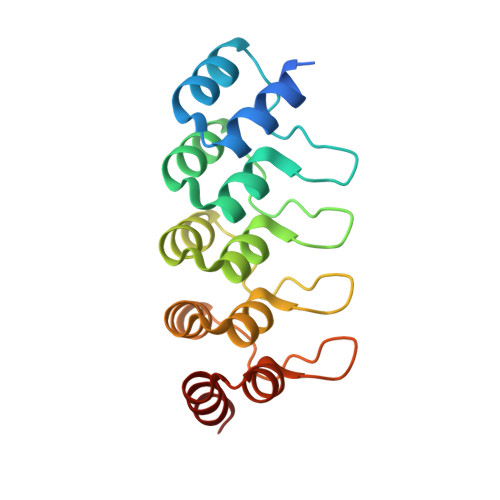
Design, construction, and characterization of a second-generation DARPin library with reduced hydrophobicity.
Seeger, M.A., Zbinden, R., Flutsch, A., Gutte, P.G., Engeler, S., Roschitzki-Voser, H., Grutter, M.G.(2013) Protein Sci 22: 1239-1257
- PubMed: 23868333
- DOI: https://doi.org/10.1002/pro.2312
- Primary Citation of Related Structures:
4J7W, 4J8Y, 4JB8 - PubMed Abstract:
Designed ankyrin repeat proteins (DARPins) are well-established binding molecules based on a highly stable nonantibody scaffold. Building on 13 crystal structures of DARPin-target complexes and stability measurements of DARPin mutants, we have generated a new DARPin library containing an extended randomized surface. To counteract the enrichment of unspecific hydrophobic binders during selections against difficult targets containing hydrophobic surfaces such as membrane proteins, the frequency of apolar residues at diversified positions was drastically reduced and substituted by an increased number of tyrosines. Ribosome display selections against two human caspases and membrane transporter AcrB yielded highly enriched pools of unique and strong DARPin binders which were mainly monomeric. We noted a prominent enrichment of tryptophan residues during binder selections. A crystal structure of a representative of this library in complex with caspase-7 visualizes the key roles of both tryptophans and tyrosines in providing target contacts. These aromatic and polar side chains thus substitute the apolar residues valine, leucine, isoleucine, methionine, and phenylalanine of the original DARPins. Our work describes biophysical and structural analyses required to extend existing binder scaffolds and simplifies an existing protocol for the assembly of highly diverse synthetic binder libraries.
Organizational Affiliation:
Department of Biochemistry, University of Zurich, 8057, Zürich, Switzerland.