Crystal Structure of the CHK1
Kang, Y.N., Stuckey, J.A., Chang, P., Russell, A.J.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Serine/threonine-protein kinase Chk1 | 279 | Homo sapiens | Mutation(s): 0 Gene Names: CHEK1, CHK1 EC: 2.7.11.1 | 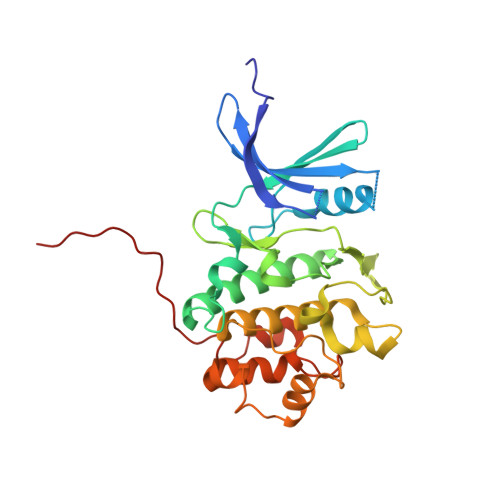 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for O14757 (Homo sapiens) Explore O14757 Go to UniProtKB: O14757 | |||||
PHAROS: O14757 GTEx: ENSG00000149554 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O14757 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HKC Query on HKC | B [auth A] | 6,7-dimethoxy-3-[4-(1H-tetrazol-5-yl)phenyl]-1,4-dihydroindeno[1,2-c]pyrazole C19 H16 N6 O2 NDAAHSGATZAMOW-UHFFFAOYSA-N |  | ||
| SO4 Query on SO4 | C [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 45.052 | α = 90 |
| b = 65.77 | β = 94.51 |
| c = 57.983 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SCALEPACK | data scaling |
| BUSTER-TNT | refinement |
| PDB_EXTRACT | data extraction |
| BUSTER | refinement |