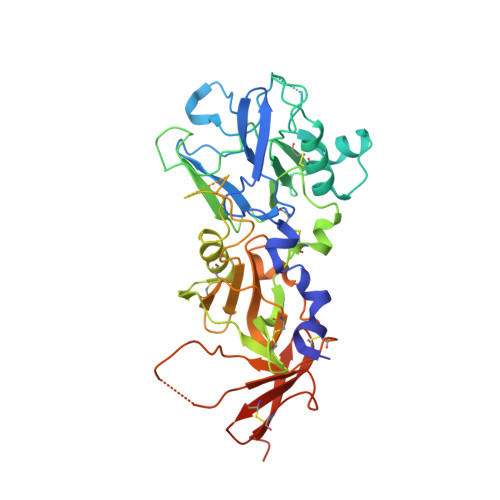
Babesia Divergens and Neospora Caninum Apical Membrane Antigen 1 (Ama1) Structures Reveal Selectivity and Plasticity in Apicomplexan Parasite Host Cell Invasion.
Tonkin, M.L., Crawford, J., Lebrun, M.L., Boulanger, M.J.(2013) Protein Sci 22: 114
- PubMed: 23169033
- DOI: https://doi.org/10.1002/pro.2193
- Primary Citation of Related Structures:
4APL, 4APM - PubMed Abstract:
Host cell invasion by the obligate intracellular apicomplexan parasites, including Plasmodium (malaria) and Toxoplasma (toxoplasmosis), requires a step-wise mechanism unique among known host-pathogen interactions. A key step is the formation of the moving junction (MJ) complex, a circumferential constriction between the apical tip of the parasite and the host cell membrane that traverses in a posterior direction to enclose the parasite in a protective vacuole essential for intracellular survival. The leading model of MJ assembly proposes that Rhoptry Neck Protein 2 (RON2) is secreted into the host cell and integrated into the membrane where it serves as the receptor for apical membrane antigen 1 (AMA1) on the parasite surface. We have previously demonstrated that the AMA1-RON2 interaction is an effective target for inhibiting apicomplexan invasion. To better understand the AMA1-dependant molecular recognition events that promote invasion, including the significant AMA1-RON2 interaction, we present the structural characterization of AMA1 from the apicomplexan parasites Babesia divergens (BdAMA1) and Neospora caninum (NcAMA1) by X-ray crystallography. These studies offer intriguing structural insight into the RON2-binding surface groove in the AMA1 apical domain, which shows clear evidence for receptor-ligand co-evolution, and the hyper variability of the membrane proximal domain, which in Plasmodium is responsible for direct binding to erythrocytes. By incorporating the structural analysis of BdAMA1 and NcAMA1 with existing AMA1 structures and complexes we were able to define conserved pockets in the AMA1 apical groove that could be targeted for the design of broadly reactive therapeutics.
Organizational Affiliation:
Department of Biochemistry & Microbiology, University of Victoria, Victoria, British Columbia, V8W 3P6, Canada.