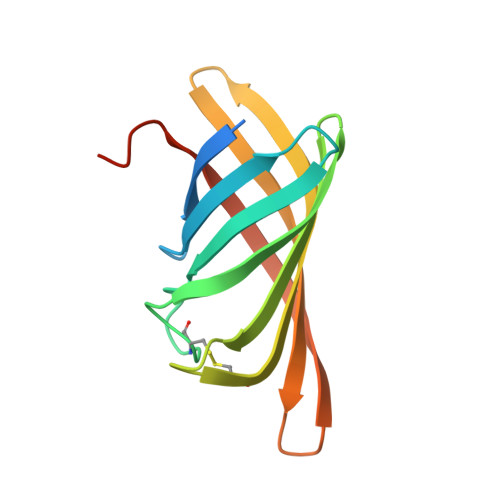
Crystal structure of rhizavidin: insights into the enigmatic high-affinity interaction of an innate biotin-binding protein dimer.
Meir, A., Helppolainen, S.H., Podoly, E., Nordlund, H.R., Hytonen, V.P., Maatta, J.A., Wilchek, M., Bayer, E.A., Kulomaa, M.S., Livnah, O.(2009) J Mol Biol 386: 379-390
- PubMed: 19111749
- DOI: https://doi.org/10.1016/j.jmb.2008.11.061
- Primary Citation of Related Structures:
3EW1, 3EW2 - PubMed Abstract:
Rhizavidin, from the proteobacterium Rhizobium etli, exhibits high affinity towards biotin but maintains an inherent dimeric quaternary structure and thus, differs from all other known tetrameric avidins. Rhizavidin also differs from the other avidins, since it lacks the characteristic tryptophan residue positioned in the L7,8 loop that plays a crucial role in high-affinity binding and oligomeric stability of the tetrameric avidins. The question is, therefore, how does the dimer exist and how is the high biotin-binding affinity retained? For this purpose, the crystal structures of apo- and biotin-complexed rhizavidin were determined. The structures reveal that the rhizavidin monomer exhibits a topology similar to those of other members of the avidin family, that is, eight antiparallel beta-strands that form the conventional avidin beta-barrel. The quaternary structure comprises the sandwich-like dimer, in which the extensive 1-4 intermonomer interface is intact, but the 1-2 and 1-3 interfaces are nonexistent. Consequently, the biotin-binding site is partially accessible, due to the lack of the tryptophan "lid" that distinguishes the tetrameric structures. In rhizavidin, a disulfide bridge connecting the L3,4 and L5,6 loops restrains the L3,4 loop conformation, leaving the binding-site residues essentially unchanged upon biotin binding. Our study suggests that in addition to the characteristic hydrogen bonding and hydrophobic interactions, the preformed architecture of the binding site and consequent shape complementarity play a decisive role in the high-affinity biotin binding of rhizavidin. The structural description of a novel dimeric avidin-like molecule will greatly contribute to the design of improved and unique avidin derivatives for diversifying the capabilities of avidin-biotin technology.
Organizational Affiliation:
Department of Biological Chemistry, The Institute of Life Sciences, The Wolfson Centre for Applied Structural Biology, The Hebrew University of Jerusalem, Givat Ram, Israel.