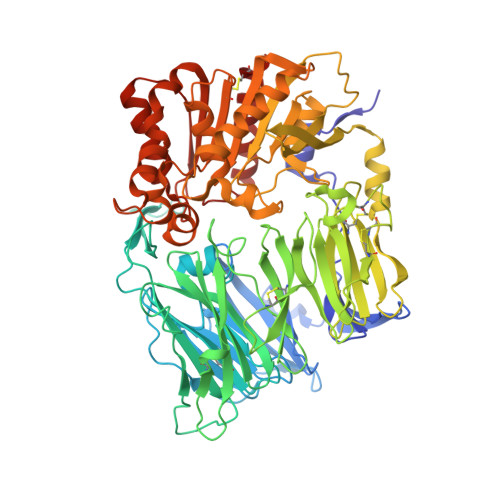
Synthesis and biological evaluation of homopiperazine derivatives with beta-aminoacyl group as dipeptidyl peptidase IV inhibitors
Ahn, J.H., Park, W.S., Jun, M.A., Shin, M.S., Kang, S.K., Kim, K.Y., Rhee, S.D., Bae, M.A., Kim, K.R., Kim, S.G., Kim, S.Y., Sohn, S.K., Kang, N.S., Lee, J.O., Lee, D.H., Cheon, H.G., Kim, S.S.(2008) Bioorg Med Chem Lett 18: 6525-6529
- PubMed: 18996694
- DOI: https://doi.org/10.1016/j.bmcl.2008.10.076
- Primary Citation of Related Structures:
3EIO - PubMed Abstract:
Compounds with homopiperazine skeleton are designed to find a potent DPP-IV inhibitor without inhibiting CYP. Thus a series of beta-aminoacyl-containing homopiperazine derivatives was synthesized and evaluated. Compounds with acid moiety were found to be potent inhibitors of DPP-IV without inhibiting CYP 3A4. More specifically, compound 7m showed nanomolar activity with no inhibition towards five subtypes of CYPs, was considered as a prototype for further derivatization. Based on its X-ray co-crystal structure with human DPP-IV, we identified compounds 7s and 7t which showed good in vitro activity, no CYP inhibition, and good selectivity.
Organizational Affiliation:
Drug Discovery Division, Korea Research Institute of Chemical Technology, Yuseong-Gu, Daejeon 305-600, Republic of Korea. jhahn@krict.re.kr