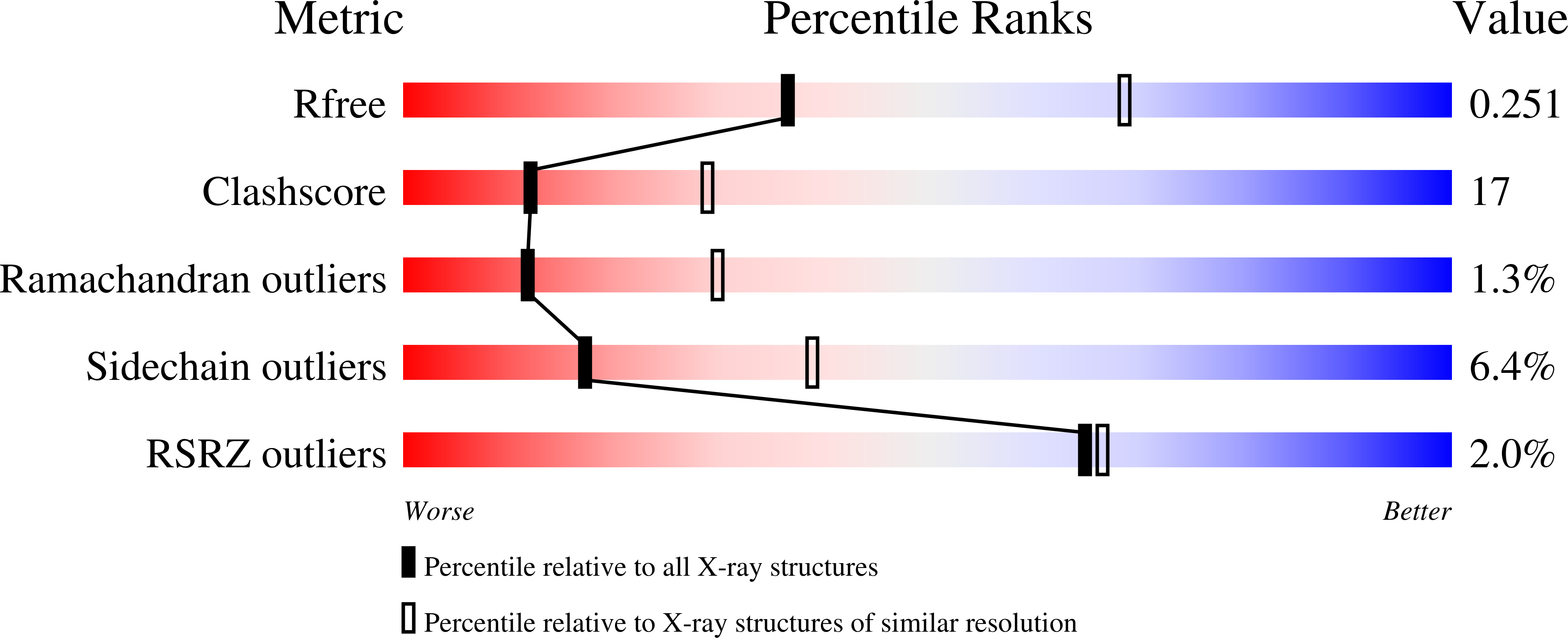
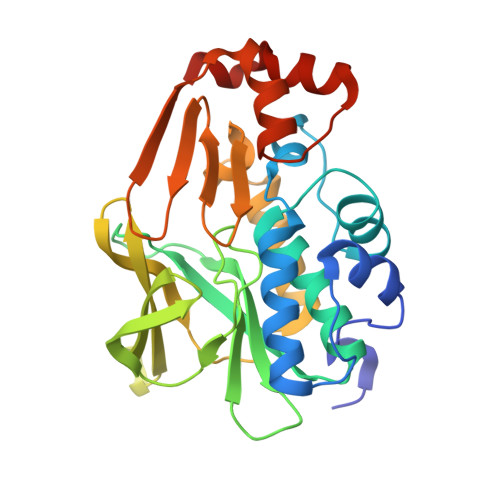
Functional and structural characterization of the arylamine N-acetyltransferase from the opportunistic pathogen Nocardia farcinica
Martins, M., Pluvinage, B., Li de la Sierra-Gallay, I., Barbault, F., Dairou, J., Dupret, J.M., Rodrigues-Lima, F.(2008) J Mol Biol 383: 549-560
- PubMed: 18778714
- DOI: https://doi.org/10.1016/j.jmb.2008.08.035
- Primary Citation of Related Structures:
3D9W - PubMed Abstract:
Arylamine N-acetyltransferase (NAT) enzymes are found in a broad range of eukaryotes and prokaryotes. There is increasing evidence that NAT enzymes could contribute to antibiotic resistance in pathogenic bacteria such as Mycobacterium tuberculosis. Nocardia farcinica is an opportunistic human pathogen that causes pulmonary infections (nocardiosis) with clinical manifestations that resemble tuberculosis. While the genomic sequence of this prokaryote has been determined, studies of N. farcinica proteins remain almost nonexistent. In particular, N. farcinica proteins putatively involved in antibiotic resistance mechanisms have not been described structurally or functionally. Here, we have characterized a new NAT enzyme (NfNAT) from N. farcinica at the structural and functional level. NfNAT is the first N. farcinica protein for which a 3D structure is reported. We showed that this novel prokaryotic isoform is structurally and functionally related to the mycobacterial NAT enzymes. In particular, NfNAT was found to display high N-acetyltransferase activity towards several known NAT substrates including the antitubercular drug isoniazid. Interestingly, isoniazid is not used for the treatment of nocardiosis and has been shown to be poorly active against several nocardial species. On the contrary, NfNAT was found to be poorly active towards sulfamethoxazole, a sulfonamide drug considered as the treatment of choice for the treatment of nocardiosis. The functional and structural data reported in this study will help to develop our understanding of the role of NAT enzymes in nocardia and mycobacteria and may help in the rational design of NAT antagonists for a range of clinical applications.
Organizational Affiliation:
Laboratoire de Cytophysiologie et Toxicologie Cellulaire (EA 1553), Université Paris Diderot-Paris 7, 75005 Paris, France.