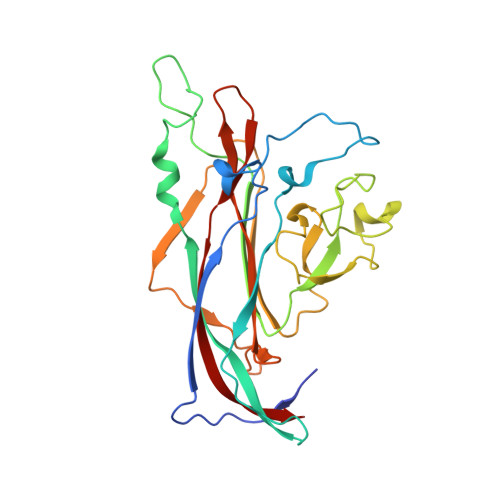
Structural basis of GM1 ganglioside recognition by simian virus 40.
Neu, U., Woellner, K., Gauglitz, G., Stehle, T.(2008) Proc Natl Acad Sci U S A 105: 5219-5224
- PubMed: 18353982
- DOI: https://doi.org/10.1073/pnas.0710301105
- Primary Citation of Related Structures:
3BWQ, 3BWR - PubMed Abstract:
Simian virus 40 (SV40) has been a paradigm for understanding attachment and entry of nonenveloped viruses, viral DNA replication, and virus assembly, as well as for endocytosis pathways associated with caveolin and cholesterol. We find by glycan array screening that SV40 recognizes its ganglioside receptor GM1 with a quite narrow specificity, but isothermal titration calorimetry shows that individual binding sites have a relatively low affinity, with a millimolar dissociation constant. The high-resolution crystal structure of recombinantly produced SV40 capsid protein, VP1, in complex with the carbohydrate portion of GM1, reveals that the receptor is bound in a shallow solvent-exposed groove at the outer surface of the capsid. Through a complex network of interactions, VP1 recognizes a conformation of GM1 that is the dominant one in solution. Analysis of contacts provides a structural basis for the observed specificity and suggests binding mechanisms for additional physiologically relevant GM1 variants. Comparison with murine Polyomavirus (Polyoma) receptor complexes reveals that SV40 uses a different mechanism of sialic acid binding, which has implications for receptor binding of human polyomaviruses. The SV40-GM1 complex reveals a parallel to cholera toxin, which uses a similar cell entry pathway and binds GM1 in the same conformation.
Organizational Affiliation:
Interfaculty Institute for Biochemistry, University of Tübingen, Hoppe-Seyler-Strasse 4, D-72076 Tübingen, Germany.