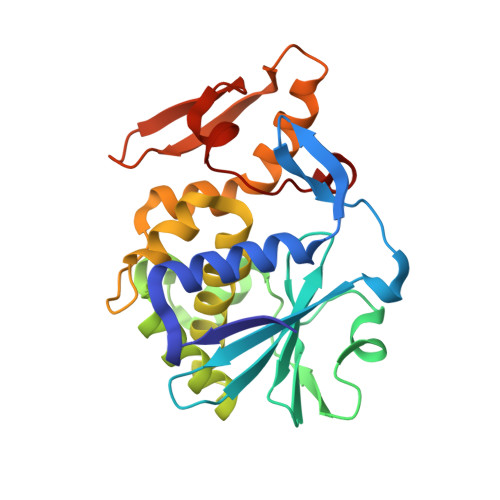
Atomic resolution structure of cucurmosin, a novel type 1 ribosome-inactivating protein from the sarcocarp of Cucurbita moschata.
Hou, X., Meehan, E.J., Xie, J., Huang, M., Chen, M., Chen, L.(2008) J Struct Biol 164: 81-87
- PubMed: 18652900
- DOI: https://doi.org/10.1016/j.jsb.2008.06.011
- Primary Citation of Related Structures:
3BWH - PubMed Abstract:
A novel type 1 ribosome-inactivating protein (RIP) designated cucurmosin was isolated from the sarcocarp of Cucurbita moschata (pumpkin). Besides rRNA N-glycosidase activity, cucurmosin exhibits strong cytotoxicities to three cancer cell lines of both human and murine origins, but low toxicity to normal cells. Plant genomic DNA extracted from the tender leaves was amplified by PCR between primers based on the N-terminal sequence and X-ray sequence of the C-terminal. The complete mature protein sequence was obtained from N-terminal protein sequencing and partial DNA sequencing, confirmed by high resolution crystal structure analysis. The crystal structure of cucurmosin has been determined at 1.04A, a resolution that has never been achieved before for any RIP. The structure contains two domains: a large N-terminal domain composed of seven alpha-helices and eight beta-strands, and a smaller C-terminal domain consisting of three alpha-helices and two beta-strands. The high resolution structure established a glycosylation pattern of GlcNAc(2)Man(3)Xyl. Asn225 was identified as a glycosylation site. Residues Tyr70, Tyr109, Glu158 and Arg161 define the active site of cucurmosin as an RNA N-glycosidase. The structural basis of cytotoxicity difference between cucurmosin and trichosanthin is discussed.
Organizational Affiliation:
State Key Laboratory of Structural Chemistry, Fujian Institute of Research on the Structure of Matter, The Chinese Academy of Sciences, Fuzhou, Fujian, China.