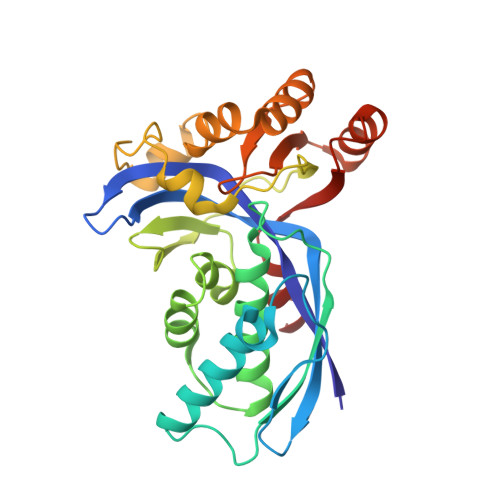
A Triclinic Crystal Form of Escherichia Coli 4-Diphosphocytidyl-2C-Methyl-D-Erythritol Kinase and Reassessment of the Quaternary Structure.
Kalinowska-Tluscik, J., Miallau, L., Gabrielsen, M., Leonard, G.A., Mcsweeney, S.M., Hunter, W.N.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 237
- PubMed: 20208151
- DOI: https://doi.org/10.1107/S1744309109054591
- Primary Citation of Related Structures:
2WW4 - PubMed Abstract:
4-Diphosphocytidyl-2C-methyl-D-erythritol kinase (IspE; EC 2.7.1.148) contributes to the 1-deoxy-D-xylulose 5-phosphate or mevalonate-independent biosynthetic pathway that produces the isomers isopentenyl diphosphate and dimethylallyl diphosphate. These five-carbon compounds are the fundamental building blocks for the biosynthesis of isoprenoids. The mevalonate-independent pathway does not occur in humans, but is present and has been shown to be essential in many dangerous pathogens, i.e. Plasmodium species, which cause malaria, and gram-negative bacteria. Thus, the enzymes involved in this pathway have attracted attention as potential drug targets. IspE produces 4-diphosphosphocytidyl-2C-methyl-D-erythritol 2-phosphate by ATP-dependent phosphorylation of 4-diphosphocytidyl-2C-methyl-D-erythritol. A triclinic crystal structure of the Escherichia coli IspE-ADP complex with two molecules in the asymmetric unit was determined at 2 A resolution and compared with a monoclinic crystal form of a ternary complex of E. coli IspE also with two molecules in the asymmetric unit. The molecular packing is different in the two forms. In the asymmetric unit of the triclinic crystal form the substrate-binding sites of IspE are occluded by structural elements of the partner, suggesting that the ;triclinic dimer' is an artefact of the crystal lattice. The surface area of interaction in the triclinic form is almost double that observed in the monoclinic form, implying that the dimeric assembly in the monoclinic form may also be an artifact of crystallization.
Organizational Affiliation:
Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland.