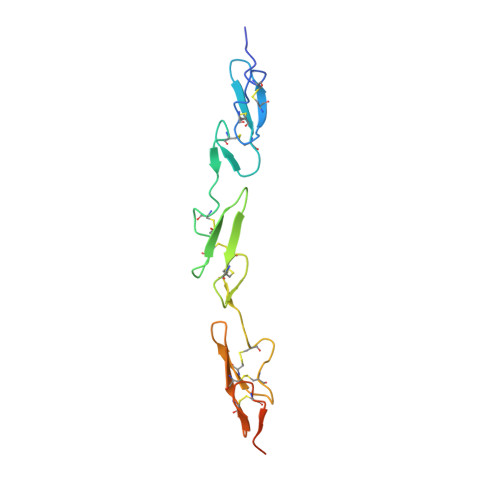
A Conserved Face of the Jagged/Serrate Dsl Domain is Involved in Notch Trans-Activation and Cis-Inhibition.
Cordle, J., Johnson, S., Tay, J.Z., Roversi, P., Wilkin, M.B., De Madrid, B.H., Shimizu, H., Jensen, S., Whiteman, P., Jin, B., Redfield, C., Baron, M., Lea, S.M., Handford, P.A.(2008) Nat Struct Mol Biol 15: 849
- PubMed: 18660822
- DOI: https://doi.org/10.1038/nsmb.1457
- Primary Citation of Related Structures:
2VJ2, 2VJ3 - PubMed Abstract:
The Notch receptor and its ligands are key components in a core metazoan signaling pathway that regulates the spatial patterning, timing and outcome of many cell-fate decisions. Ligands contain a disulfide-rich Delta/Serrate/LAG-2 (DSL) domain required for Notch trans-activation or cis-inhibition. Here we report the X-ray structure of a receptor binding region of a Notch ligand, the DSL-EGF3 domains of human Jagged-1 (J-1(DSL-EGF3)). The structure reveals a highly conserved face of the DSL domain, and we show, by functional analysis of Drosophila melanogster ligand mutants, that this surface is required for both cis- and trans-regulatory interactions with Notch. We also identify, using NMR, a surface of Notch-1 involved in J-1(DSL-EGF3) binding. Our data imply that cis- and trans-regulation may occur through the formation of structurally distinct complexes that, unexpectedly, involve the same surfaces on both ligand and receptor.
Organizational Affiliation:
Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, UK.