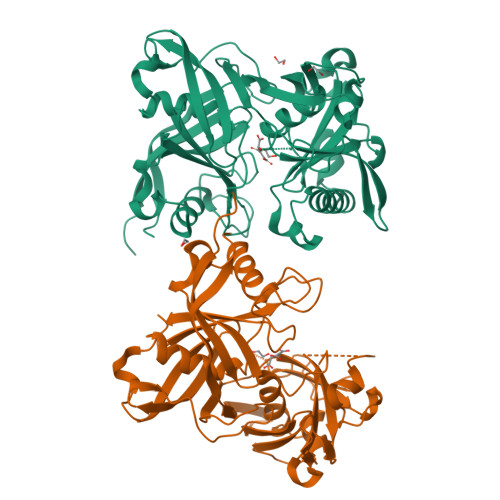
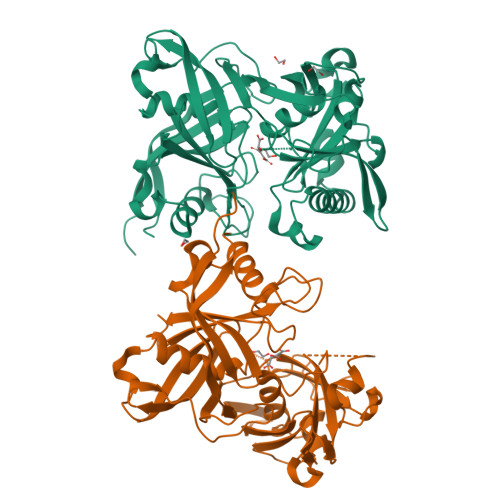
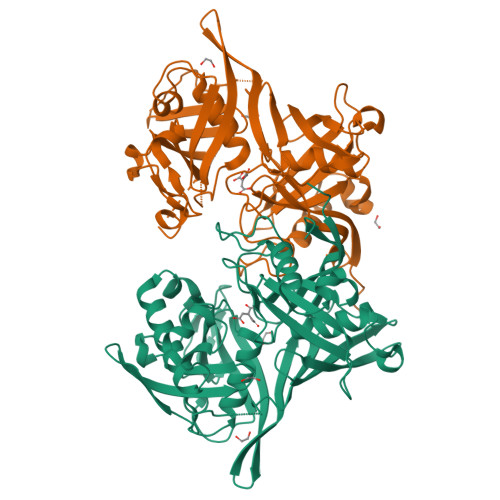
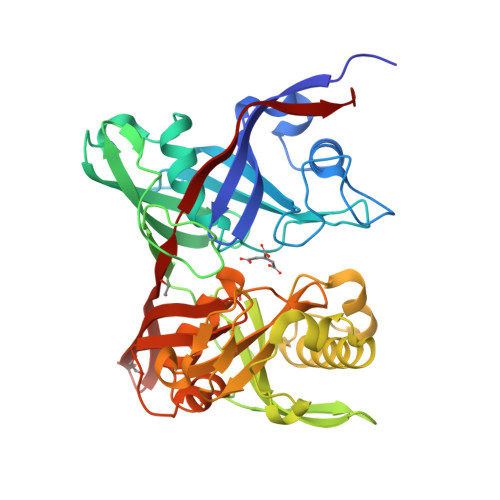
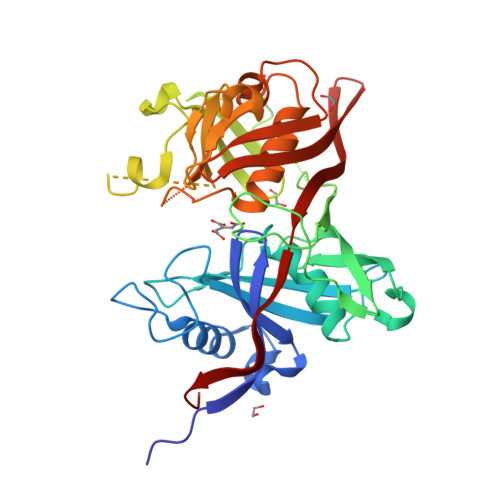
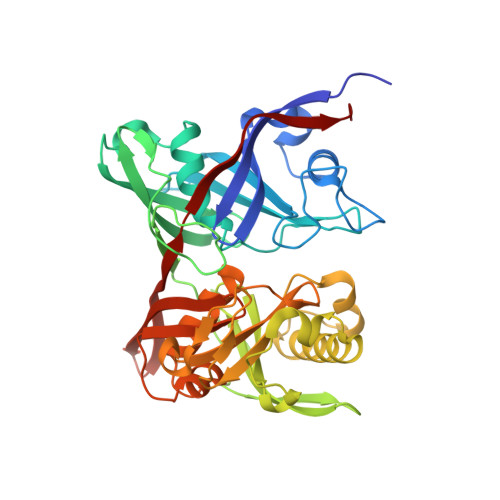
The three-dimensional crystal structure of the PrpF protein of Shewanella oneidensis complexed with trans-aconitate: insights into its biological function.
Garvey, G.S., Rocco, C.J., Escalante-Semerena, J.C., Rayment, I.(2007) Protein Sci 16: 1274-1284
- PubMed: 17567742
- DOI: https://doi.org/10.1110/ps.072801907
- Primary Citation of Related Structures:
2PVZ, 2PW0 - PubMed Abstract:
In bacteria, the dehydration of 2-methylcitrate to yield 2-methylaconitate in the 2-methylcitric acid cycle is catalyzed by a cofactor-less (PrpD) enzyme or by an aconitase-like (AcnD) enzyme. Bacteria that use AcnD also require the function of the PrpF protein, whose function was previously unknown. To gain insights into the function of PrpF, the three-dimensional crystal structure of the PrpF protein from the bacterium Shewanella oneidensis was solved at 2.0 A resolution. The protein fold of PrpF is strikingly similar to those of the non-PLP-dependent diaminopimelate epimerase from Haemophilus influenzae, a putative proline racemase from Brucella melitensis, and to a recently deposited structure of a hypothetical protein from Pseudomonas aeruginosa. Results from in vitro studies show that PrpF isomerizes trans-aconitate to cis-aconitate. It is proposed that PrpF catalysis of the cis-trans isomerization proceeds through a base-catalyzed proton abstraction coupled with a rotation about C2-C3 bond of 2-methylaconitate, and that residue Lys73 is critical for PrpF function. The newly identified function of PrpF as a non-PLP-dependent isomerase, together with the fact that PrpD-containing bacteria do not require PrpF, suggest that the isomer of 2-methylaconitate that serves as a substrate of aconitase must have the same stereochemistry as that synthesized by PrpD. From this, it follows that the 2-methylaconitate isomer generated by AcnD is not a substrate of aconitase, and that PrpF is required to generate the correct isomer. As a consequence, the isomerase activity of PrpF may now be viewed as an integral part of the 2-methylcitric acid cycle.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison 53706, USA.