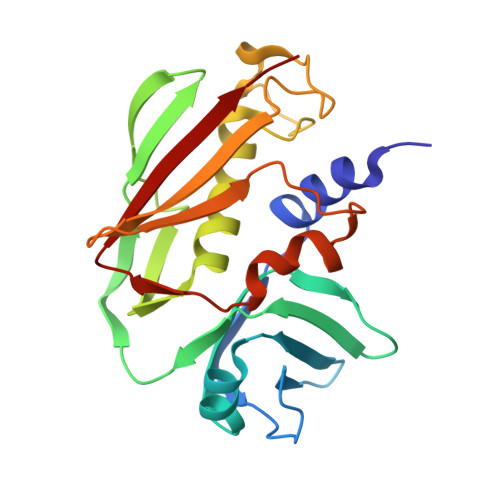
A novel loop domain in superantigens extends their T cell receptor recognition site
Gunther, S., Varma, A.K., Moza, B., Kasper, K.J., Wyatt, A.W., Zhu, P., Rahman, A.K., Li, Y., Mariuzza, R.A., McCormick, J.K., Sundberg, E.J.(2007) J Mol Biol 371: 210-221
- PubMed: 17560605
- DOI: https://doi.org/10.1016/j.jmb.2007.05.038
- Primary Citation of Related Structures:
2NTS, 2NTT - PubMed Abstract:
Superantigens (SAGs) interact with host immune receptors to induce a massive release of inflammatory cytokines that can lead to toxic shock syndrome and death. Bacterial SAGs can be classified into five distinct evolutionary groups. Group V SAGs are characterized by the alpha3-beta8 loop, a unique approximately 15 amino acid residue extension that is required for optimal T cell activation. Here, we report the X-ray crystal structures of the group V SAG staphylococcal enterotoxin K (SEK) alone and in complex with the TCR hVbeta5.1 domain. SEK adopts a unique TCR binding orientation relative to other SAG-TCR complexes, which results in the alpha3-beta8 loop contacting the apical loop of framework region 4, thereby extending the known TCR recognition site of SAGs. These interactions are absolutely required for TCR binding and T cell activation by SEK, and dictate the TCR Vbeta domain specificity of SEK and other group V SAGs.
Organizational Affiliation:
Boston Biomedical Research Institute, Watertown, MA 02472, USA.