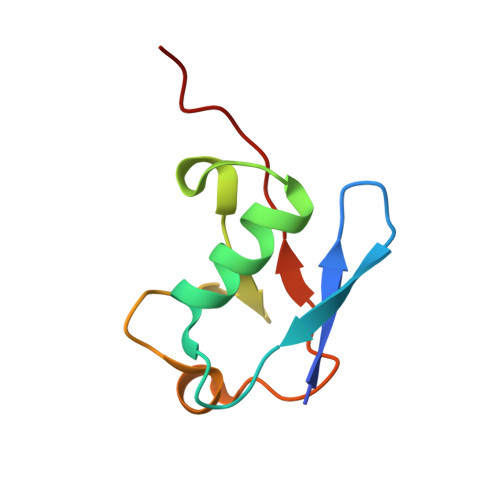
Conformational dynamics control ubiquitin-deubiquitinase interactions and influence in vivo signaling.
Phillips, A.H., Zhang, Y., Cunningham, C.N., Zhou, L., Forrest, W.F., Liu, P.S., Steffek, M., Lee, J., Tam, C., Helgason, E., Murray, J.M., Kirkpatrick, D.S., Fairbrother, W.J., Corn, J.E.(2013) Proc Natl Acad Sci U S A 110: 11379-11384
- PubMed: 23801757
- DOI: https://doi.org/10.1073/pnas.1302407110
- Primary Citation of Related Structures:
2M0X - PubMed Abstract:
Ubiquitin is a highly conserved eukaryotic protein that interacts with a diverse set of partners to act as a cellular signaling hub. Ubiquitin's conformational flexibility has been postulated to underlie its multifaceted recognition. Here we use computational and library-based means to interrogate core mutations that modulate the conformational dynamics of human ubiquitin. These ubiquitin variants exhibit increased affinity for the USP14 deubiquitinase, with concomitantly reduced affinity for other deubiquitinases. Strikingly, the kinetics of conformational motion are dramatically slowed in these variants without a detectable change in either the ground state fold or excited state population. These variants can be ligated into substrate-linked chains in vitro and in vivo but cannot solely support growth in eukaryotic cells. Proteomic analyses reveal nearly identical interaction profiles between WT ubiquitin and the variants but identify a small subset of altered interactions. Taken together, these results show that conformational dynamics are critical for ubiquitin-deubiquitinase interactions and imply that the fine tuning of motion has played a key role in the evolution of ubiquitin as a signaling hub.
Organizational Affiliation:
Department of Early Discovery Biochemistry, Genentech Inc., South San Francisco, CA 94080, USA.