Identification of structural traits that increase the antimicrobial activity of a chimeric peptide of human beta-defensins 2 and 3.
Spudy, B., Sonnichsen, F.D., Waetzig, G.H., Grotzinger, J., Jung, S.(2012) Biochem Biophys Res Commun 427: 207-211
- PubMed: 22995312
- DOI: https://doi.org/10.1016/j.bbrc.2012.09.052
- Primary Citation of Related Structures:
2LXO - PubMed Abstract:
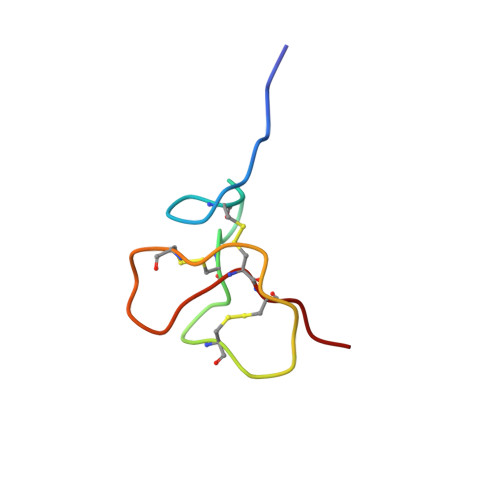
Antimicrobial peptides participate in the first line of defence of many organisms against pathogens. In humans, the family of β-defensins plays a pivotal role in innate immunity. Two human β-defensins, β-defensin-2 and -3 (HBD2 and HBD3), show substantial sequence identity and structural similarity. However, HBD3 kills Staphylococcus (S.) aureus with a 4- to 8-fold higher efficiency compared to HBD2, whereas their activities against Escherichia (E.) coli are very similar. The generation of six HBD2/HBD3-chimeric molecules led to the identification of distinct molecular regions which mediate their divergent killing properties. One of the chimeras (chimera C3) killed both E. coli and S. aureus with an even higher efficacy compared to the wild-type molecules. Due to the broad spectrum of its antimicrobial activity against many human multidrug-resistant pathogens, this HBD2/HBD3-chimeric peptide represents a promising candidate for a new class of antibiotics. In order to investigate the structural basis of its exceptional antimicrobial activity, the peptide's tertiary structure was determined by NMR spectroscopy, which allowed its direct comparison to the published structures of HBD2 and HBD3 and the identification of the activity-increasing molecular features.
Organizational Affiliation:
Institute of Biochemistry, Christian-Albrechts-University, Olshausenstr. 40, 24098 Kiel, Germany.