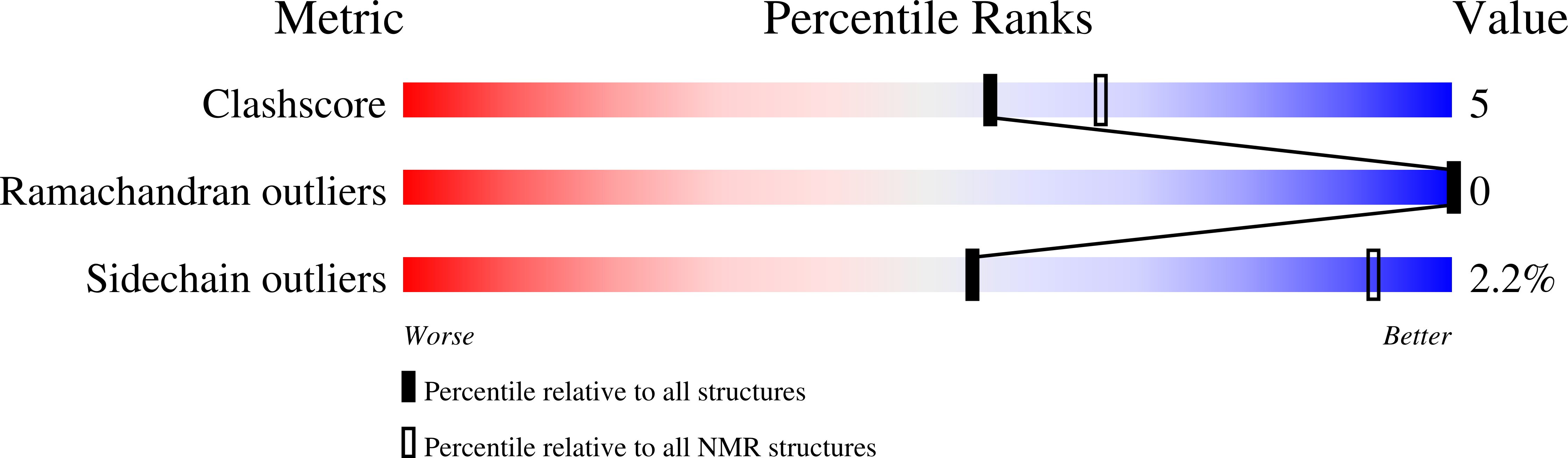An engineered second disulfide bond restricts lymphotactin/XCL1 to a chemokine-like conformation with XCR1 agonist activity
Tuinstra, R.L., Peterson, F.C., Elgin, E.S., Pelzek, A.J., Volkman, B.F.(2007) Biochemistry 46: 2564-2573
- PubMed: 17302442
- DOI: https://doi.org/10.1021/bi602365d
- Primary Citation of Related Structures:
2HDM - PubMed Abstract:
Chemokines adopt a conserved tertiary structure stabilized by two disulfide bridges and direct the migration of leukocytes. Lymphotactin (Ltn) is a unique chemokine in that it contains only one disulfide and exhibits large-scale structural heterogeneity. Under physiological solution conditions (37 degrees C and 150 mM NaCl), Ltn is in equilibrium between the canonical chemokine fold (Ltn10) and a distinct four-stranded beta-sheet (Ltn40). Consequently, it has not been possible to address the biological significance of each structural species independently. To stabilize the Ltn10 structure in a manner independent of specific solution conditions, Ltn variants containing a second disulfide bridge were designed. Placement of the new cysteines was based on a sequence alignment of Ltn with either the first (Ltn-CC1) or third disulfide (Ltn-CC3) in the CC chemokine, HCC-2. NMR data demonstrate that both CC1 and CC3 retain the Ltn10 chemokine structure and no longer exhibit structural rearrangement. The ability of each mutant to activate the Ltn receptor, XCR1, has been tested using an intracellular Ca2+ flux assay. These data support the conclusion that the chemokine fold of Ltn10 is responsible for receptor activation. We also examined the role of amino- and carboxyl-terminal residues in Ltn-mediated receptor activation. In contrast to previous reports, we find that the 25 residues comprising the novel C-terminal extension do not participate in receptor activation, while the native N-terminus is absolutely required for Ltn function.
Organizational Affiliation:
Department of Biochemistry, Medical College of Wisconsin, Milwaukee, Wisconsin 53226, USA.