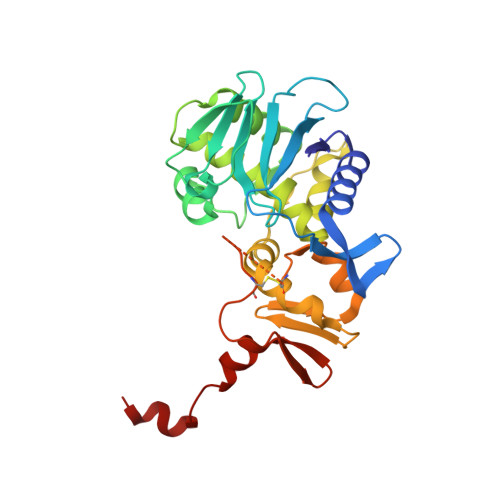
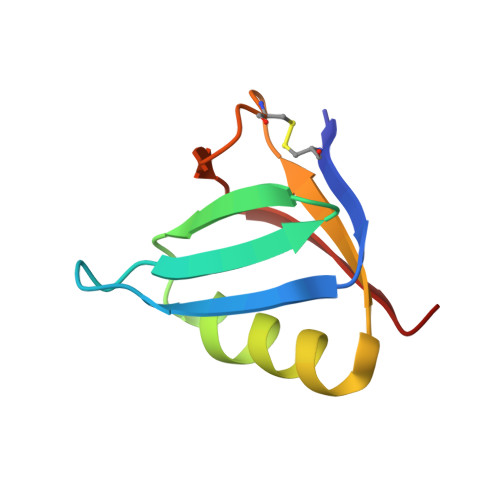
Binding of adenine to Stx2, the protein toxin from Escherichia coli O157:H7.
Fraser, M.E., Cherney, M.M., Marcato, P., Mulvey, G.L., Armstrong, G.D., James, M.N.(2006) Acta Crystallogr Sect F Struct Biol Cryst Commun 62: 627-630
- PubMed: 16820678
- DOI: https://doi.org/10.1107/S1744309106021968
- Primary Citation of Related Structures:
2GA4 - PubMed Abstract:
Stx2 is a protein toxin whose catalytic subunit acts as an N-glycosidase to depurinate a specific adenine base from 28S rRNA. In the holotoxin, the catalytic portion, A1, is linked to the rest of the A subunit, A2, and A2 interacts with the pentameric ring formed by the five B subunits. In order to test whether the holotoxin is active as an N-glycosidase, Stx2 was crystallized in the presence of adenosine and adenine. The crystals diffracted to approximately 1.8 angstroms and showed clear electron density for adenine in the active site. Adenosine had been cleaved, proving that Stx2 is an active N-glycosidase. While the holotoxin is active against small substrates, it would be expected that the B subunits would interfere with the binding of the 28S rRNA.
Organizational Affiliation:
Department of Biological Sciences, University of Calgary, 2500 University Drive NW, Calgary AB T2N 1N4, Canada. frasm@ucalgary.ca