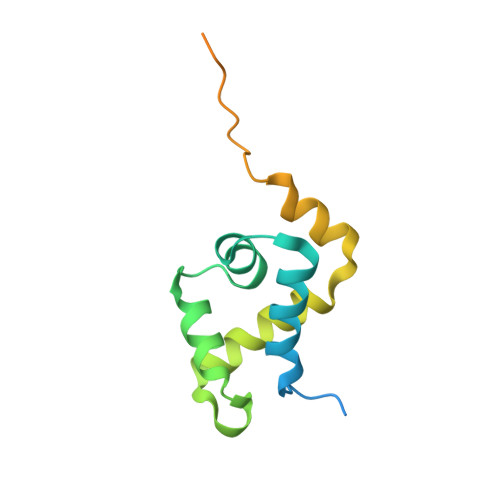
Solution Structure and DNA-binding Mode of the Matrix Attachment Region-binding Domain of the Transcription Factor SATB1 That Regulates the T-cell Maturation
Yamaguchi, H., Tateno, M., Yamasaki, K.(2006) J Biol Chem 281: 5319-5327
- PubMed: 16371359
- DOI: https://doi.org/10.1074/jbc.M510933200
- Primary Citation of Related Structures:
1YSE - PubMed Abstract:
SATB1 is a transcriptional regulator controlling the gene expression that is essential in the maturation of the immune T-cell. SATB1 binds to the nuclear matrix attachment regions of DNA, where it recruits histone deacetylase and represses transcription through a local chromatin remodeling. Here we determined the solution structure of the matrix attachment region-binding domain, possessing similarity to the CUT DNA-binding domain, of human SATB1 by NMR spectroscopy. The structure consists of five alpha-helices, in which the N-terminal four are arranged similarly to the four-helix structure of the CUT domain of hepatocyte nuclear factor 6alpha. By an NMR chemical shift perturbation analysis and by surface plasmon resonance analyses of SATB1 mutant proteins, an interface for DNA binding was revealed to be located at the third helix and the surrounding regions. Surface plasmon resonance experiments using groove-specific binding drugs and methylated DNAs indicated that the domain recognizes DNA from the major groove side. These observations suggested that SATB1 possesses a DNA-binding mode similar to that of the POU-specific DNA-binding domain, which is known to share structural similarity to the four-helix CUT domain.
Organizational Affiliation:
Age Dimension Research Center, National Institute of Advanced Industrial Science and Technology, 1-1-1 Higashi, Tsukuba 305-8566, Japan.