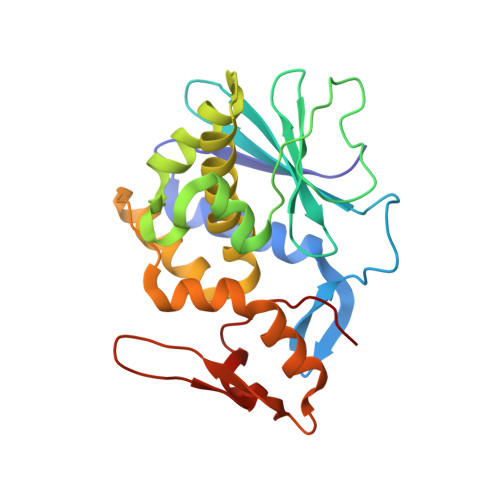
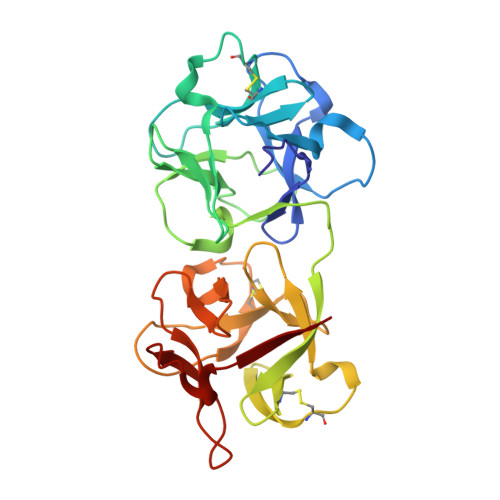
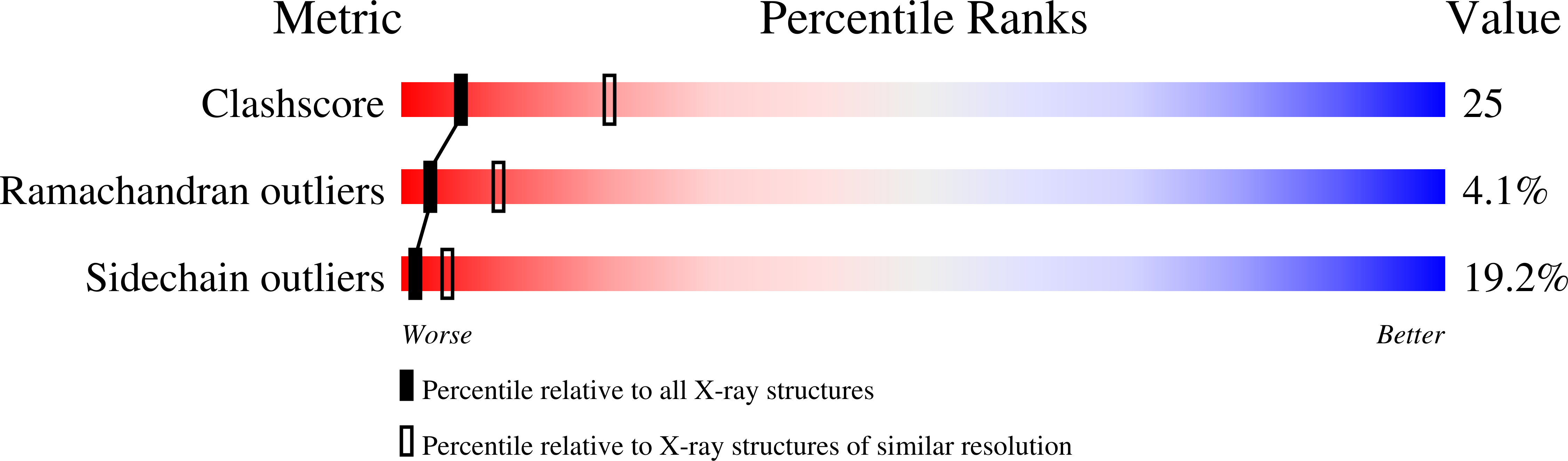Crystal structure of himalayan mistletoe ribosome-inactivating protein reveals the presence of a natural inhibitor and a new functionally active sugar-binding site.
Mishra, V., Bilgrami, S., Sharma, R.S., Kaur, P., Yadav, S., Krauspenhaar, R., Betzel, C., Voelter, W., Babu, C.R., Singh, T.P.(2005) J Biol Chem 280: 20712-20721
- PubMed: 15774467
- DOI: https://doi.org/10.1074/jbc.M500735200
- Primary Citation of Related Structures:
1YF8 - PubMed Abstract:
Ribosome-inactivating proteins (RIPs) are toxins involved in plant defense. How the plant prevents autotoxicity is not yet fully understood. The present study is the first structural evidence of a naturally inhibited form of RIP from a plant. Himalayan mistletoe RIP (HmRIP) was purified from Viscum album leaves and crystallized with lactose. The structure was determined by the molecular replacement method and refined at 2.8-A resolution. The crystal structure revealed the presence of high quality non-protein electron density at the active site, into which a pteridine derivative (2-amino 4-isopropyl 6-carboxyl pteridine) was modeled. The carboxyl group of the ligand binds strongly with the key active site residue Arg(162), nullifies the positive charge required for catalysis, and thereby acts as a natural inhibitor. Lectin subunits of RIPs have two active sugar-binding sites present in 1alpha- and 2gamma-subdomains. A third functionally active site has been identified in the 1beta-subdomain of HmRIP. The 1beta-site is active despite the absence of conserved polar sugar-binding residues. Loss of these residues is compensated by the following: (i) the presence of an extended site where the penultimate sugar also interacts with the protein; (ii) the interactions of galactose with the protein main chain carbonyl and amide nitrogen atoms; (iii) the presence of a well defined pocket encircled by four walls; and (iv) a favorable stacking of the galactose ring with Tyr(66) besides the conserved Phe(75). The mode of sugar binding is also distinct at the 1alpha and 2gamma sugar-binding sites.
Organizational Affiliation:
Department of Biophysics, All India Institute of Medical Sciences, New Delhi 110 029, India.