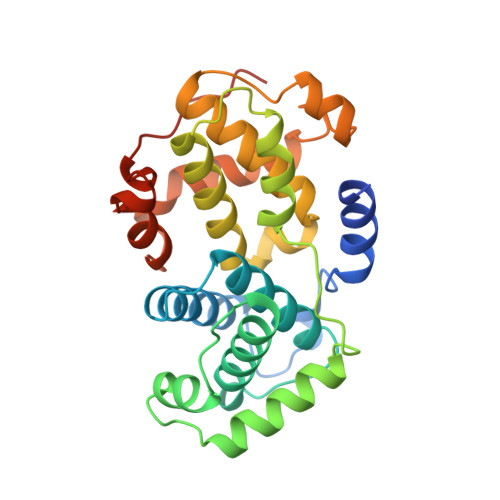
The crystal structure of cyclin A.
Brown, N.R., Noble, M.E., Endicott, J.A., Garman, E.F., Wakatsuki, S., Mitchell, E., Rasmussen, B., Hunt, T., Johnson, L.N.(1995) Structure 3: 1235-1247
- PubMed: 8591034
- DOI: https://doi.org/10.1016/s0969-2126(01)00259-3
- Primary Citation of Related Structures:
1VIN - PubMed Abstract:
Eukaryotic cell cycle progression is regulated by cyclin dependent protein kinases (CDKs) whose activity is regulated by association with cyclins and by reversible phosphorylation. Cyclins also determine the subcellular location and substrate specificity of CDKs. Cyclins exhibit diverse sequences but all share homology over a region of approximately 100 amino acids, termed the cyclin box. From the determination of the structure of cyclin A, together with results from biochemical and genetic analyses, we can identify which parts of the cyclin molecular may contribute to cyclin A structure and function. We have solved the crystal structure, at 2.0 A resolution, of an active recombinant fragment of bovine cyclin A, cyclin A-3, corresponding to residues 171-432 of human cyclin A. The cyclin box has an alpha-helical fold comprising five alpha helices. This fold is repeated in the C-terminal region, although this region shares negligible sequence similarity with the cyclin box. Analysis of residues that are conserved throughout the A, B, and E cyclins identifies two exposed clusters of residues, one of which has recently been shown to be involved in the association with human CDK2. The second cluster may identify another site of cyclin A-protein interaction. Comparison of the structure of the unbound cyclin with the structure of cyclin A complexed with CDK2 reveals that cyclin A does not undergo any significant conformational changes on complex formation. Threading analysis shows that the cyclin-box fold is consistent with the sequences of the transcription factor TFIIB and other functionally related proteins. The structural results indicate a role for the cyclin-box fold both as a template for the cyclin family and as a generalised adaptor molecule in the regulation of transcription.
Organizational Affiliation:
Laboratory of Molecular Biophysics, Oxford, UK.