NMR Solution Structure of the Archaebacterial Chromosomal Protein MC1 Reveals a New Protein Fold
Paquet, F., Culard, F., Barbault, F., Maurizot, J.C., Lancelot, G.(2004) Biochemistry 43: 14971-14978
- PubMed: 15554704
- DOI: https://doi.org/10.1021/bi048382z
- Primary Citation of Related Structures:
1T23 - PubMed Abstract:
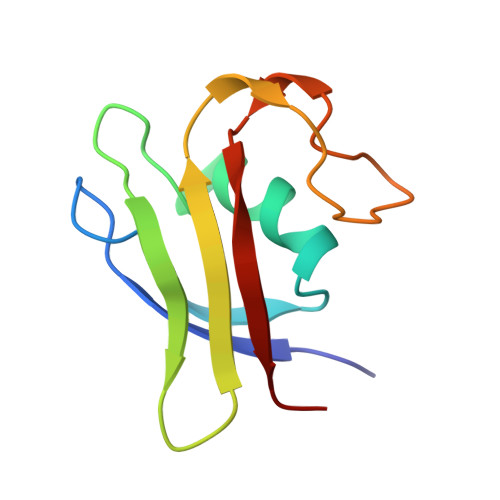
The three-dimensional structure of methanogen chromosomal protein 1 (MC1), a chromosomal protein extracted from the archaebacterium Methanosarcina sp. CHTI55, has been solved using (1)H NMR spectroscopy. The small basic protein MC1 contains 93 amino acids (24 basic residues against 12 acidic residues). The main elements of secondary structures are an alpha helix and five beta strands, arranged as two antiparallel beta sheets (a double one and a triple one) packed in an orthogonal manner forming a barrel. The protein displays a largely hydrophilic surface and a very compact hydrophobic core made up by side chains at the interface of the two beta sheets and the helix side facing the interior of the protein. The MC1 solution structure shows a globular protein with overall dimensions in the range of 34-40 A, which potentially corresponds to a DNA-binding site of 10-12 base pairs. The presumed DNA-binding site is located on the sequence comprising residues K62-P82, which is formed by a part of strands II2 and II3 belonging to the triple-stranded antiparallel beta sheet and a loop flanked by prolines P68 and P76. The tryptophan W74 that is expected to play a key role in the DNA-binding according to photocross-linking experiments was found completely exposed to the solvent, in a good position to interact with DNA. The overall fold of MC1, characterized by its linking beta-beta-alpha-beta-beta-loop-beta, is different from other known DNA-binding proteins. Its structure suggests a different DNA-binding mode than those of the histone-like proteins HU or HMGB. Thus, MC1 may be classified as a member of a new family.
Organizational Affiliation:
Centre de Biophysique Moléculaire, CNRS, Rue Charles Sadron, 45071 Orléans Cedex 2, France. paquet@cnrs-orleans.fr