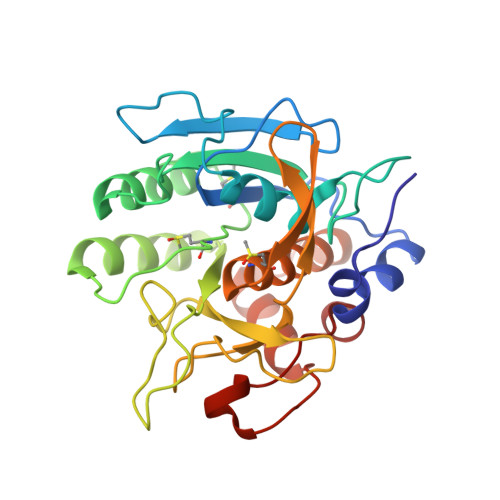
The three-dimensional structure of Bacillus amyloliquefaciens subtilisin at 1.8 A and an analysis of the structural consequences of peroxide inactivation.
Bott, R., Ultsch, M., Kossiakoff, A., Graycar, T., Katz, B., Power, S.(1988) J Biol Chem 263: 7895-7906
- PubMed: 3286644
- Primary Citation of Related Structures:
1ST2, 2ST1 - PubMed Abstract:
The three-dimensional structure of the subtilisin from Bacillus amyloliquefaciens (BAS) has been refined to 1.8 A using the amino acid sequence deduced from the DNA coding sequence. The structure is essentially the same as the previously reported structures of subtilisin BPN' (Wright, C.S., Alden, R.A., and Kraut, J. (1969) Nature 221, 235-242) and Novo (Drenth, J., Hol, W. G. J., Jansonius, J. N., and Koekoek, R. (1972) Eur. J. Biochem. 26, 177-181) determined in different crystal forms, at 2.5 and 2.8 A resolution, respectively. The largest differences in the three crystallographic models are seen in regions where the amino acid sequence used in the fit to the electron density maps of BPN' and Novo differs from the gene sequence of BAS (Wells, J. A., Ferrari, E., Henner, D. J., Estell, D. A., and Chen, E. Y. (1983) Nucleic Acids Res. 11, 7911-7925). The refined BAS model shows new features of cation binding, hydrogen bonding, and internal solvent structure. The refined BAS model has served as a basis for the analysis of stereochemical factors involved in the peroxide inactivation of the enzyme. Methionine 222, which is adjacent to the catalytic Ser221, is quantitatively oxidized to the sulfoxide by hydrogen peroxide as had been previously shown for the related Bacillus licheniformis enzyme (Stauffer, C. E., and Etson, D. (1969) J. Biol. Chem. 244, 5333-5338). In addition to this site of modification, we observe partial to full oxidation of two of the four remaining methionines. The oxidation of the methionines does not correlate well with their solvent accessibility calculated from the x-ray structure coordinates; in addition, only one of the two possible stereoisomers of methionine sulfoxide is formed. We also detect hydrogen peroxide-induced modification of the hydroxyl groups of two tyrosines. Modeling suggests that most of the observed effect of oxidation on the enzyme's catalytic efficiency can be attributed to unfavorable interactions at the oxyanion binding site between the sulfoxide group at 222 and the carbonyl oxygen of the scissile peptide bond of the bound substrate.
Organizational Affiliation:
Department of Biomolecular Chemistry, Genentech, Inc., South San Francisco, California 94080.