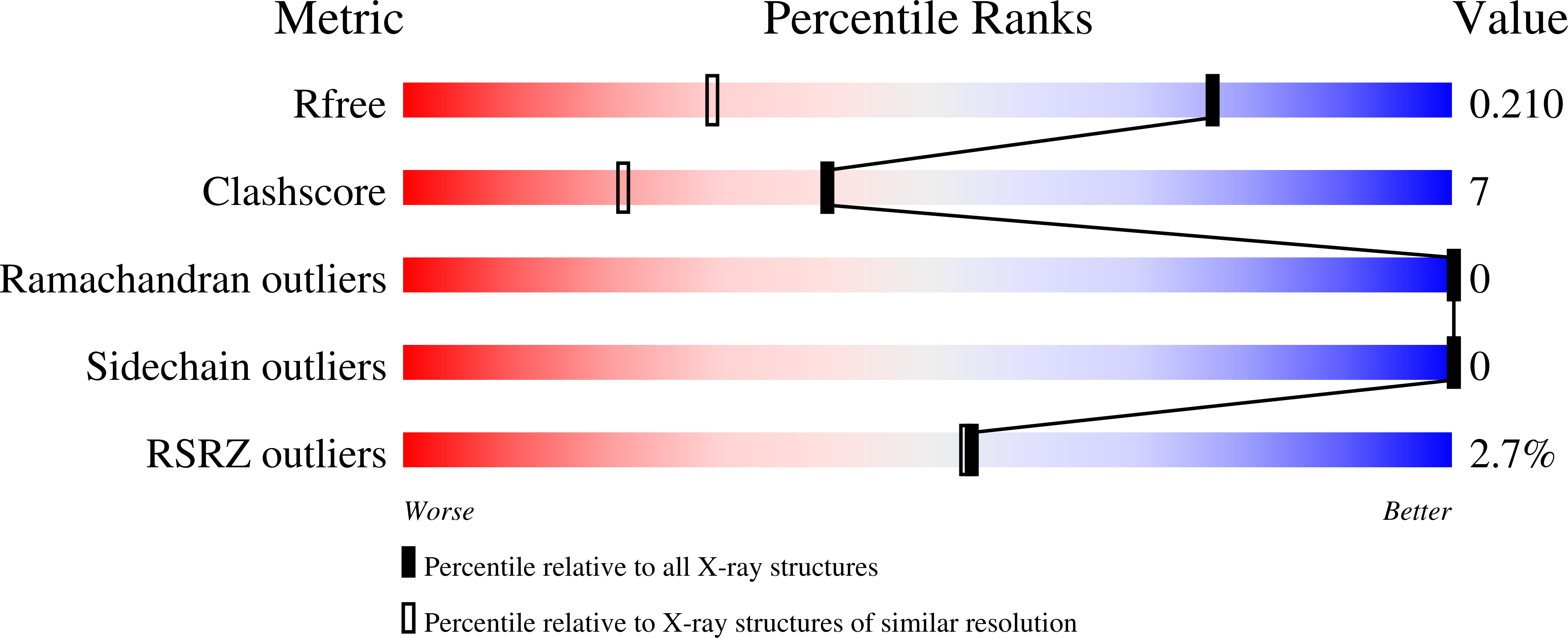
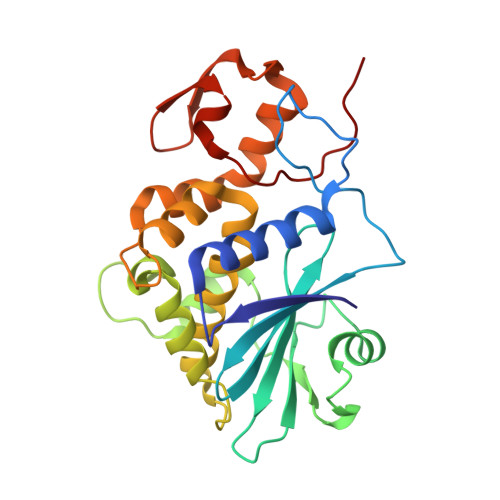
The 1.4A structure of dianthin 30 indicates a role of surface potential at the active site of type 1 ribosome inactivating proteins
Fermani, S., Falini, G., Ripamonti, A., Polito, L., Stirpe, F., Bolognesi, A.(2005) J Struct Biol 149: 204-212
- PubMed: 15681236
- DOI: https://doi.org/10.1016/j.jsb.2004.11.007
- Primary Citation of Related Structures:
1RL0 - PubMed Abstract:
Ribosome inactivating proteins (RIPs) are plant proteins with enzymatic activity identified as rRNA N-glycosidase (EC 3.2.2.22), which cleaves the N-glycosidic bond of a specific adenine on the ricin/sarcin region of rRNA, thus causing inhibition of protein synthesis. They also depurinate extensively DNA and other polynucleotides. The three-dimensional structure of dianthin 30, a type 1 (single-chain) RIP of Dianthus caryophyllus (leaves), is now described at 1.4 angstroms, a resolution never achieved before for any RIP. The fold typical of RIPs is conserved, despite some differences in the loop regions. The general structure comparison by superimposed alpha-carbon (249 atoms) and the sequence alignment by structure for dianthin 30 and saporin-S6 give a root mean square deviation of 0.625 angstroms. Despite the differences reported for the biological activities of the two RIPs, their structures fit quite well and both show a protein segment containing strands beta7, beta8, and beta9 shorter than other RIPs. However, the surface electrostatic potential in the active site region neatly distinguishes dianthin 30 from saporin-S6. The possible relationship between the charge distribution and the behavior of the proteins toward different substrates is discussed.
Organizational Affiliation:
Dipartimento di Chimica G. Ciamician, Alma Mater Studiorum Universita' di Bologna, via Selmi 2, I-40126 Bologna, Italy.