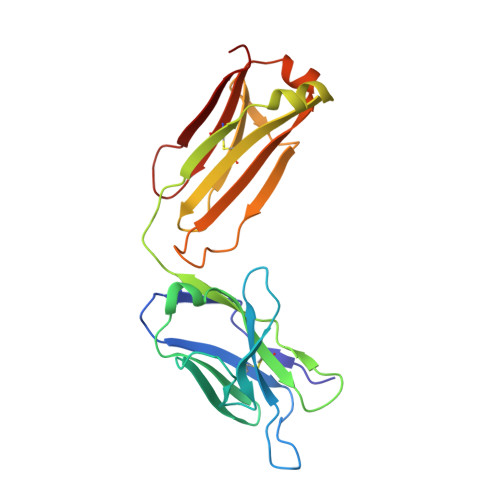
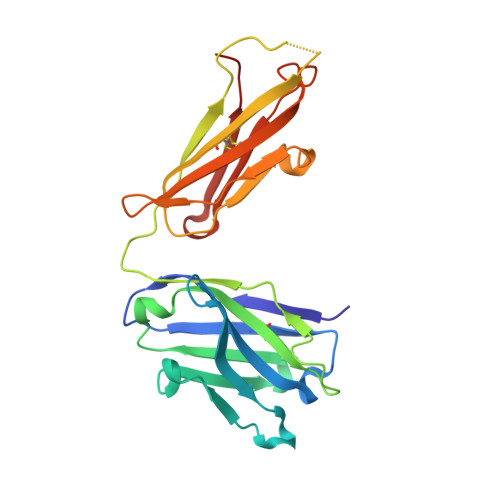
Diversity in hapten recognition: structural study of an anti-cocaine antibody M82G2.
Pozharski, E., Moulin, A., Hewagama, A., Shanafelt, A.B., Petsko, G.A., Ringe, D.(2005) J Mol Biol 349: 570-582
- PubMed: 15885702
- DOI: https://doi.org/10.1016/j.jmb.2005.03.080
- Primary Citation of Related Structures:
1Q72, 1QYG, 1RFD - PubMed Abstract:
Antibodies against cocaine and other drugs of abuse are the basis for diagnostic tests for the presence of those drugs in human serum. The 1.7A resolution crystal structure of the anti-cocaine monoclonal antibody M82G2 in complex with cocaine is presented. This structure determination was undertaken to establish the stereochemical features in the antibody binding site that confer specificity for cocaine, and as part of an ongoing project to understand the rules that govern molecular recognition. The cocaine-binding site can be characterized topologically as a narrow groove on the protein surface. The antibody utilizes water-mediated hydrogen bonding, and cation-pi and stacking (pi-pi) interactions to provide specificity. Comparison with the previously published structure of the anti-cocaine antibody GNC92H2 shows that binding of a small ligand can be achieved in diverse ways, both in terms of a binding site structure/topology and protein-ligand interactions.
Organizational Affiliation:
Rosenstiel Basic Medical Sciences Research Center, Brandeis University, 415 South Street, Waltham, MA 02454, USA.