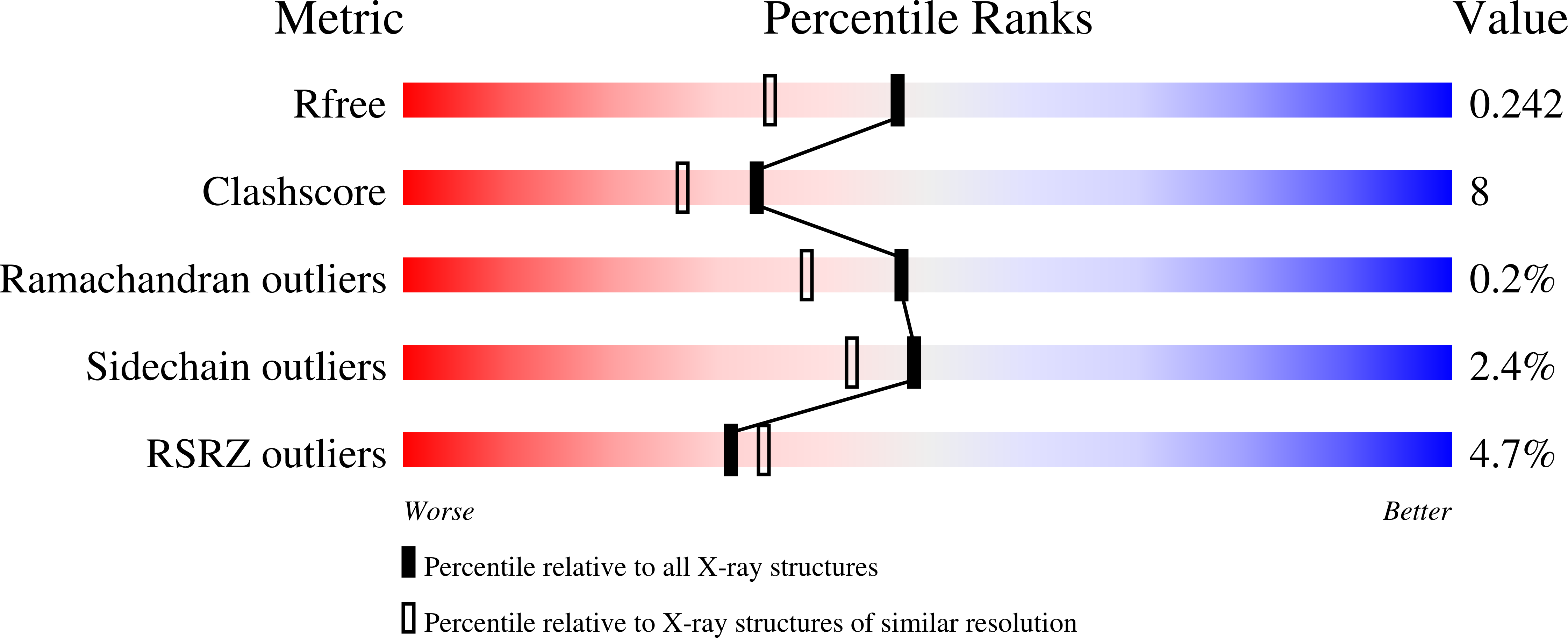
STRUCTURAL CHARACTERIZATION OF A HUMAN CYTOSOLIC NMN/NaMN ADENYLYLTRANSFERASE AND IMPLICATION IN HUMAN NAD BIOSYNTHESIS
ZHANG, X., KURNASOV, O.V., KARTHIKEYAN, S., GRISHIN, N.V., OSTERMAN, A.L., ZHANG, H.(2003) J Biol Chem 278: 13503-13511
- PubMed: 12574164
- DOI: https://doi.org/10.1074/jbc.M300073200
- Primary Citation of Related Structures:
1NUP, 1NUQ, 1NUR, 1NUS, 1NUT, 1NUU - PubMed Abstract:
Pyridine dinucleotides (NAD and NADP) are ubiquitous cofactors involved in hundreds of redox reactions essential for the energy transduction and metabolism in all living cells. In addition, NAD also serves as a substrate for ADP-ribosylation of a number of nuclear proteins, for silent information regulator 2 (Sir2)-like histone deacetylase that is involved in gene silencing regulation, and for cyclic ADP ribose (cADPR)-dependent Ca(2+) signaling. Pyridine nucleotide adenylyltransferase (PNAT) is an indispensable central enzyme in the NAD biosynthesis pathways catalyzing the condensation of pyridine mononucleotide (NMN or NaMN) with the AMP moiety of ATP to form NAD (or NaAD). Here we report the identification and structural characterization of a novel human PNAT (hsPNAT-3) that is located in the cytoplasm and mitochondria. Its subcellular localization and tissue distribution are distinct from the previously identified human nuclear PNAT-1 and PNAT-2. Detailed structural analysis of PNAT-3 in its apo form and in complex with its substrate(s) or product revealed the catalytic mechanism of the enzyme. The characterization of the cytosolic human PNAT-3 provided compelling evidence that the final steps of NAD biosynthesis pathways may exist in mammalian cytoplasm and mitochondria, potentially contributing to their NAD/NADP pool.
Organizational Affiliation:
Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, Texas 75390, USA. zhang@chop.swmed.edu