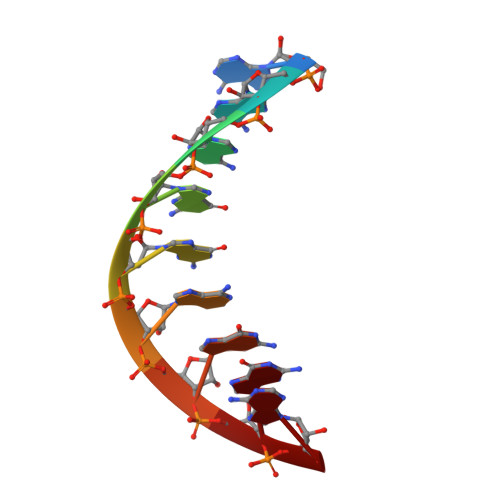
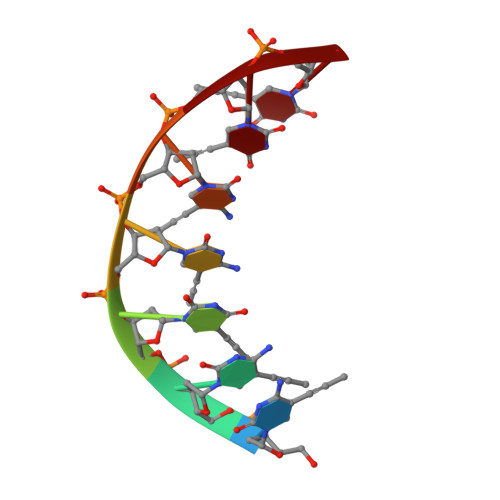
NMR Studies of DNA Single Strands and DNA:RNA Hybrids With and Without 1-Propynylation at C5 of Oligopyrimidines
Znosko, B.M., Barnes III, T.W., Krugh, T.R., Turner, D.H.(2003) J Am Chem Soc 125: 6090-6097
- PubMed: 12785839
- DOI: https://doi.org/10.1021/ja021285d
- Primary Citation of Related Structures:
1NTQ, 1NTS, 1NTT - PubMed Abstract:
The 1-propynylation at C5 of consecutive pyrimidines in DNA can enhance DNA:RNA hybrid stability at 37 degrees C by over 1 kcal/mol of substitution [Barnes, T. W., III; Turner, D. H. J. Am. Chem. Soc.2001, 123, 4107-4118]. To provide information on the structural consequences of propynylation, two-dimensional NMR spectroscopy was used to study the structures of several oligonucleotides. Intraresidue nuclear Overhauser effect spectroscopy cross peaks were observed at 30 degrees C and a 200 ms mixing time in the H6-H1' region for 5'(dC(P)C(P)U(P)C(P)C(P)U(P)U(P)) (ssPrODN) but not for 5'(dCCUCCUU) (ssODN), suggesting preorganization of the propynylated single strand. NMR structures of the duplexes 5'(dC(P)C(P)U(P)C(P)C(P)U(P)U(P))3':3'(rGAGGAGGAAAU)5' (PrODN:RNA), 5'(dCC(P)U(P)C(P)C(P)U(P)U(P))3':3'(rGAGGAGGAAAU)5' (sPrODN1:RNA), and 5'(dCCUCCUU)3':3'(rGAGGAGGAAAU)5' (ODN:RNA) indicate that their global structures are almost identical. The NMR data, however, suggest that the 5'-end of sPrODN1:RNA is more dynamic than that of PrODN:RNA. In the propynylated duplexes, the propyne group stacks on the aromatic ring of the 5'-base and extends into the major groove. The results suggest that the increased stability of the propynylated duplexes is caused by preorganization of the propynylated single strand and different interactions in the double strand. The propynyl group provides volume exclusion, enhanced stacking, and possibly different solvation.
Organizational Affiliation:
Department of Chemistry, University of Rochester, New York 14627-0216, USA.