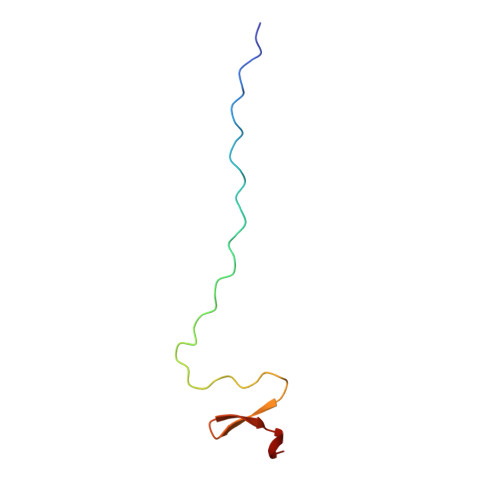
Collagen Stabilization at Atomic Level. Crystal Structure of Designed (GlyProPro)(10)foldon
Stetefeld, J., Frank, S., Jenny, M., Schulthess, T., Kammerer, R.A., Boudko, S., Landwehr, R., Okuyama, K., Engel, J.(2003) Structure 11: 339-346
- PubMed: 12623021
- DOI: https://doi.org/10.1016/s0969-2126(03)00025-x
- Primary Citation of Related Structures:
1NAY - PubMed Abstract:
In a designed fusion protein the trimeric domain foldon from bacteriophage T4 fibritin was connected to the C terminus of the collagen model peptide (GlyProPro)(10) by a short Gly-Ser linker to facilitate formation of the three-stranded collagen triple helix. Crystal structure analysis at 2.6 A resolution revealed conformational changes within the interface of both domains compared with the structure of the isolated molecules. A striking feature is an angle of 62.5 degrees between the symmetry axis of the foldon trimer and the axis of the triple helix. The melting temperature of (GlyProPro)(10) in the designed fusion protein (GlyProPro)(10)foldon is higher than that of isolated (GlyProPro)(10,) which suggests an entropic stabilization compensating for the destabilization at the interface.
Organizational Affiliation:
Department of Biophysical Chemistry, University of Basel, Klingelbergstrasse 70, CH-4056, Basel, Switzerland. joerg.stetefeld@unibas.ch