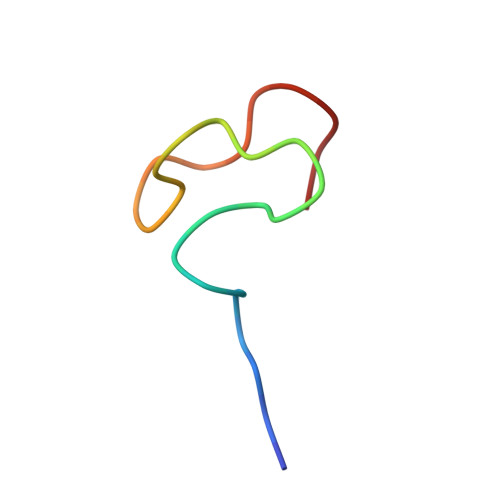
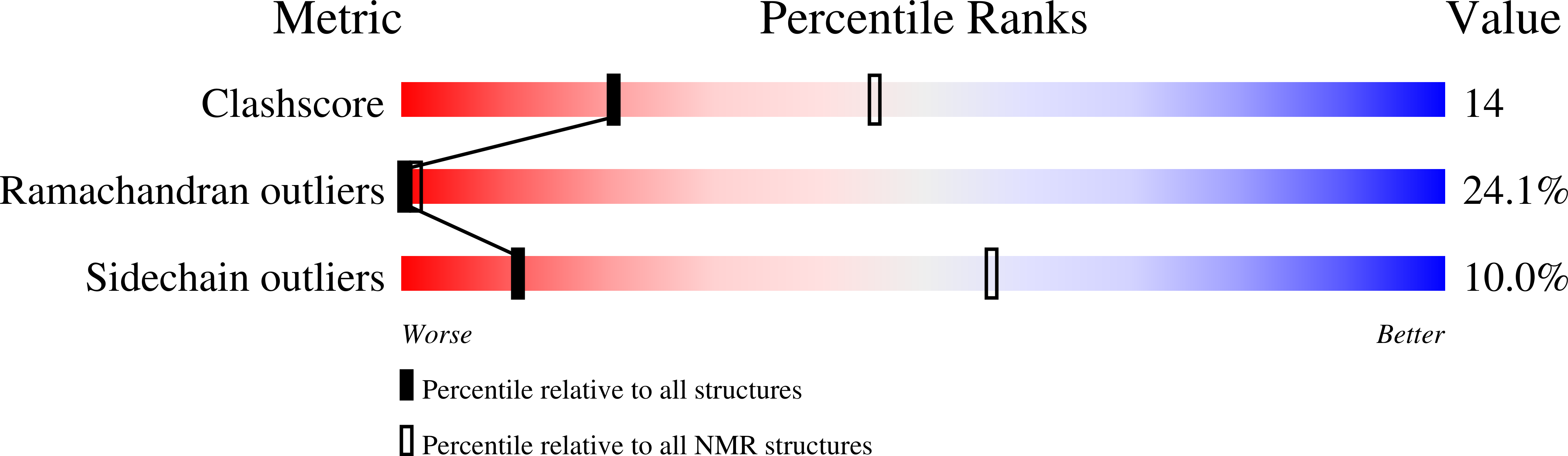Methionyl-tRNA synthetase zinc binding domain. Three-dimensional structure and homology with rubredoxin and gag retroviral proteins.
Fourmy, D., Dardel, F., Blanquet, S.(1993) J Mol Biol 231: 1078-1089
- PubMed: 8515466
- DOI: https://doi.org/10.1006/jmbi.1993.1353
- Primary Citation of Related Structures:
1MEA, 1MED - PubMed Abstract:
Methionyl-tRNA synthetase from Escherichia coli contains one tightly bound zinc atom per subunit. The region encompassing residues 138 to 163 of this enzyme is responsible for the metal binding. A 28-mer peptide corresponding to these residues was expressed in vivo and shown to contain approximately 1 mol of tightly bound Zn/mol of peptide. In this study, the three-dimensional solution structure of this peptide was solved by means of two-dimensional proton NMR spectroscopy. A total of 133 nuclear Overhauser effect distance constraints and 22 dihedral angle restraints were used for the calculations, using a hybrid distance-geometry-simulated annealing strategy. Excluding the first four residues, the resulting structure is well-defined (r.m.s.d. 0.71 A for backbone atoms) and composed of a series of four tight turns. The second and the fourth turns are composed of CXXC sequences which are structurally homologous to the NH-S turns found in the metal binding sites of gag retroviral proteins and rubredoxin. The solution structure of the zinc binding peptide shows significant discrepancies with the crystal structure of methionyl-tRNA synthetase.
Organizational Affiliation:
Laboratoire de Biochimie, URA 240 CNRS Ecole Polytechnique, Palaiseau, France.