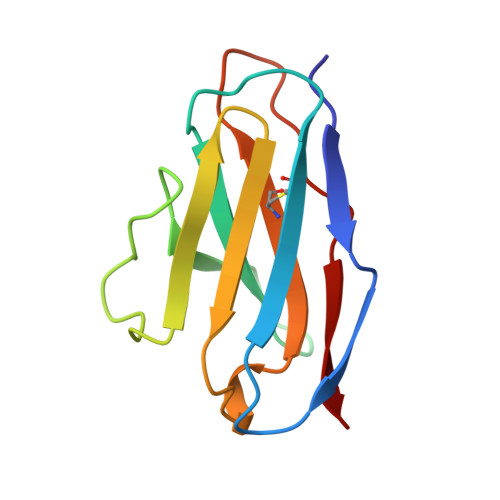
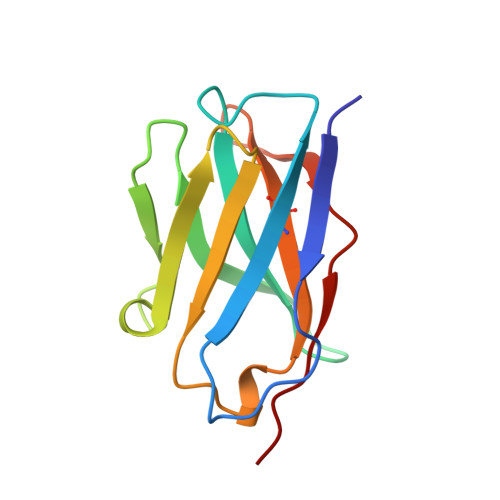
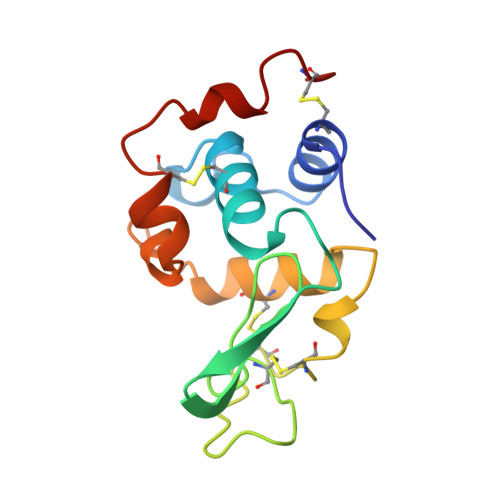
Hydrogen bonding and solvent structure in an antigen-antibody interface. Crystal structures and thermodynamic characterization of three Fv mutants complexed with lysozyme.
Fields, B.A., Goldbaum, F.A., Dall'Acqua, W., Malchiodi, E.L., Cauerhff, A., Schwarz, F.P., Ysern, X., Poljak, R.J., Mariuzza, R.A.(1996) Biochemistry 35: 15494-15503
- PubMed: 8952503
- DOI: https://doi.org/10.1021/bi961709e
- Primary Citation of Related Structures:
1KIP, 1KIQ, 1KIR - PubMed Abstract:
Using site-directed mutagenesis, X-ray crystallography, and titration calorimetry, we have examined the structural and thermodynamic consequences of removing specific hydrogen bonds in an antigen-antibody interface. Crystal structures of three antibody FvD1.3 mutants, VLTyr50Ser (VLY50S), VHTyr32Ala (VHY32A), and VHTyr101Phe (VHY101F), bound to hen egg white lysozyme (HEL) have been determined at resolutions ranging from 1.85 to 2.10 A. In the wild-type (WT) FvD1.3-HEL complex, the hydroxyl groups of VLTyr50, VHTyr32, and VHTyr101 each form at least one hydrogen bond with the lysozyme antigen. Thermodynamic parameters for antibody-antigen association have been measured using isothermal titration calorimetry, giving equilibrium binding constants Kb (M-1) of 2.6 x 10(7) (VLY50S), 7.0 x 10(7) (VHY32A), and 4.0 x 10(6) (VHY101F). For the WT complex, Kb is 2.7 x 10(8) M-1; thus, the affinities of the mutant Fv fragments for HEL are 10-, 4-, and 70-fold lower than that of the original antibody, respectively. In all three cases entropy compensation results in an affinity loss that would otherwise be larger. Comparison of the three mutant crystal structures with the WT structure demonstrates that the removal of direct antigen-antibody hydrogen bonds results in minimal shifts in the positions of the remaining protein atoms. These observations show that this complex is considerably tolerant, both structurally and thermodynamically, to the truncation of antibody side chains that form hydrogen bonds with the antigen. Alterations in interface solvent structure for two of the mutant complexes (VLY50S and VHY32A) appear to compensate for the unfavorable enthalpy changes when protein-protein interactions are removed. These changes in solvent structure, along with the increased mobility of side chains near the mutation site, probably contribute to the observed entropy compensation. For the VHY101F complex, the nature of the large entropy compensation is not evident from a structural comparison of the WT and mutant complexes. Differences in the local structure and dynamics of the uncomplexed Fv molecules may account for the entropic discrepancy in this case.
Organizational Affiliation:
Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, Rockville, Maryland 20850, USA.