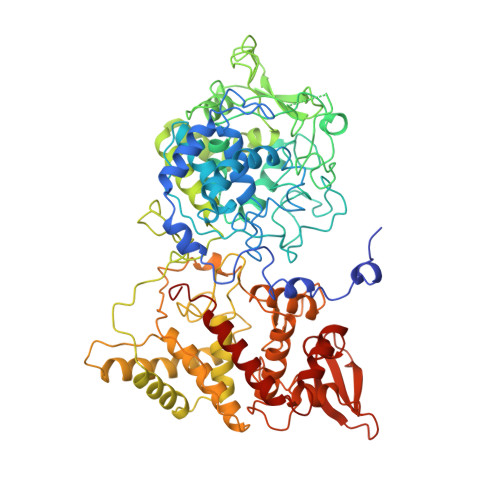
The 2.0 A crystal structure of catalase-peroxidase from Haloarcula marismortui.
Yamada, Y., Fujiwara, T., Sato, T., Igarashi, N., Tanaka, N.(2002) Nat Struct Biol 9: 691-695
- PubMed: 12172540
- DOI: https://doi.org/10.1038/nsb834
- Primary Citation of Related Structures:
1ITK - PubMed Abstract:
Catalase-peroxidase is a member of the class I peroxidase superfamily. The enzyme exhibits both catalase and peroxidase activities to remove the harmful peroxide molecule from the living cell. The 2.0 A crystal structure of the catalase-peroxidase from Haloarcula marismortui (HmCP) reveals that the enzyme is a dimer of two identical subunits. Each subunit is composed of two structurally homologous domains with a topology similar to that of class I peroxidase. The active site of HmCP is in the N-terminal domain. Although the arrangement of the catalytic residues and the cofactor heme b in the active site is virtually identical to that of class I peroxidases, the heme moiety is buried inside the domain, similar to that in a typical catalase. In the vicinity of the active site, novel covalent bonds are formed among the side chains of three residues, including that of a tryptophan on the distal side of the heme. Together with the C-terminal domain, these covalent bonds fix two long loops on the surface of the enzyme that cover the substrate access channel to the active site. These features provide an explanation for the dual activities of this enzyme.
Organizational Affiliation:
Department of Life Science, Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, 4259 Nagatsuta, Midori-ku, Yokohama 226-8501, Japan.