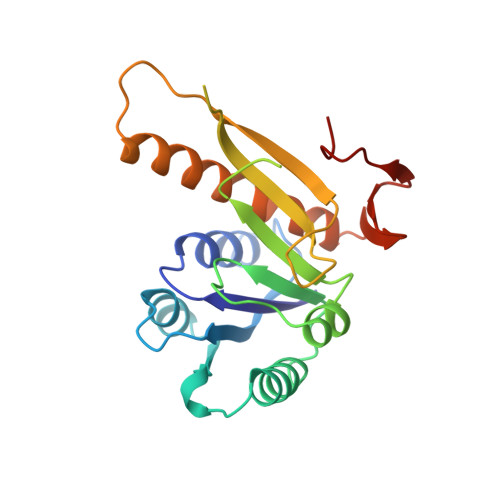
Crystal structure of glycinamide ribonucleotide transformylase from Escherichia coli at 3.0 A resolution. A target enzyme for chemotherapy.
Chen, P., Schulze-Gahmen, U., Stura, E.A., Inglese, J., Johnson, D.L., Marolewski, A., Benkovic, S.J., Wilson, I.A.(1992) J Mol Biol 227: 283-292
- PubMed: 1522592
- DOI: https://doi.org/10.1016/0022-2836(92)90698-j
- Primary Citation of Related Structures:
1GRC - PubMed Abstract:
The atomic structure of glycinamide ribonucleotide transformylase, an essential enzyme in purine biosynthesis, has been determined at 3.0 A resolution. The last three C-terminal residues and a sequence stretch of 18 residues (residues 113 to 130) are not visible in the electron density map. The enzyme forms a dimer in the crystal structure. Each monomer is divided into two domains, which are connected by a central mainly parallel seven-stranded beta-sheet. The N-terminal domain contains a Rossmann type mononucleotide fold with a phosphate ion bound to the C-terminal end of the first beta-strand. A long narrow cleft stretches from the phosphate to a conserved aspartic acid, Asp144, which has been suggested as an active-site residue. The cleft is lined by a cluster of residues, which are conserved between bacterial, yeast, avian and human enzymes, and likely represents the binding pocket and active site of the enzyme. GAR Tfase binds a reduced folate cofactor and glycinamide ribonucleotide for the catalysis of one of the initial steps in purine biosynthesis. Folate analogs and multi-substrate inhibitors of the enzyme have antineoplastic effects and the structure determination of the unliganded enzyme and enzyme-inhibitor complexes will aid the development of anti-cancer drugs.
Organizational Affiliation:
Department of Molecular Biology, Scripps Research Institute, La Jolla, CA 92037.