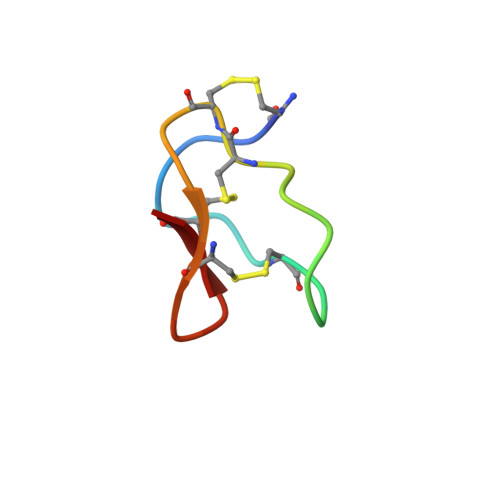
Solution structure and backbone dynamics of an omega-conotoxin precursor
Goldenberg, D.P., Koehn, R.E., Gilbert, D.E., Wagner, G.(2001) Protein Sci 10: 538-550
- PubMed: 11344322
- DOI: https://doi.org/10.1110/ps.30701
- Primary Citation of Related Structures:
1FEO - PubMed Abstract:
Nuclear magnetic resonance spectroscopy was used to characterize the solution structure and backbone dynamics of a putative precursor form of omega-conotoxin MVIIA, a 25-amino-acid residue peptide antagonist of voltage-gated Ca(2+) channels. The mature peptide is found in the venom of a fish-hunting marine snail Conus magus and contains an amidated carboxyl terminus that is generated by oxidative cleavage of a Gly residue. The form examined in this study is identical to the mature peptide except for the presence of the unmodified carboxy-terminal Gly. This form, referred to as omega-MVIIA-Gly, has previously been shown to refold and form its disulfides more efficiently than the mature form, suggesting that the presence of the terminal Gly may favor folding in vivo. The nuclear magnetic resonance (NMR) structure determination indicated that the fold of omega-MVIIA-Gly is very similar to that previously determined for the mature form, but revealed that the terminal Gly residue participates in a network of hydrogen bonds involving both backbone and side chain atoms, very likely accounting for the enhanced stability and folding efficiency. (15)N relaxation experiments indicated that the backbone is well ordered on the nanosecond time scale but that residues 9-15 undergo a conformational exchange processes with a time constant of approximately 35 microseconds. Other studies have implicated this segment in the binding of the peptide to its physiological target, and the observed motions may play a role in allowing the peptide to enter the binding site
Organizational Affiliation:
Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115, USA. goldenberg@biology.utah.edu