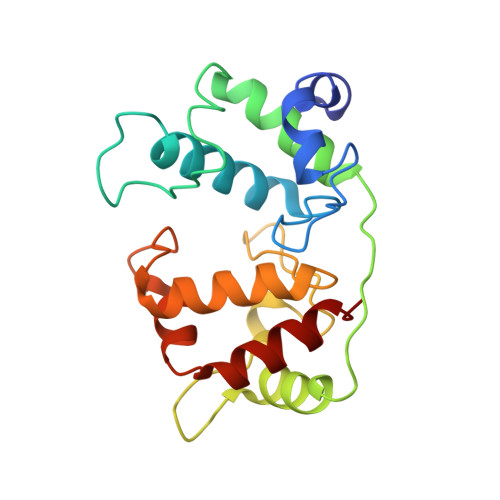
Crystal structure of the dihaem cytochrome c4 from Pseudomonas stutzeri determined at 2.2A resolution.
Kadziola, A., Larsen, S.(1997) Structure 5: 203-216
- PubMed: 9032080
- DOI: https://doi.org/10.1016/s0969-2126(97)00179-2
- Primary Citation of Related Structures:
1ETP - PubMed Abstract:
. Cytochromes c4 are dihaem cytochromes c found in a variety of bacteria. They are assumed to take part in the electron-transport systems associated with both aerobic and anaerobic respiration. The cytochrome c4 proteins are located in the periplasm, predominantly bound to the inner membrane, and are able to transfer electrons between membrane-bound reduction systems and terminal oxidases. Alignment of cytochrome c4 sequences from three bacteria, Pseudomonas aeruginosa, Pseudomonas stutzeri and Azotobacter vinelandii, suggests that these dihaem proteins are composed of two similar domains. Two distinctly different redox potentials have been measured for the Ps. stutzeri cytochrome c4, however.
Organizational Affiliation:
Centre for Crystallographic Studies, Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen, Denmark.