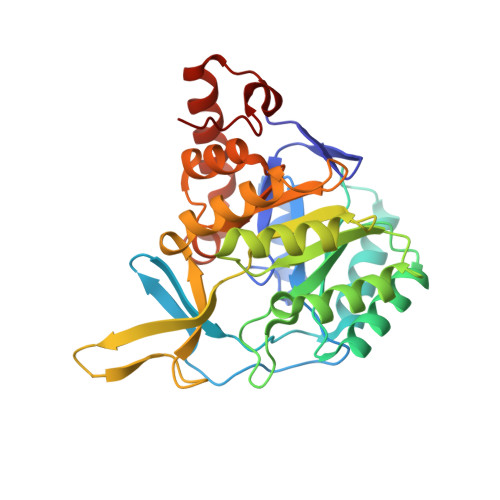
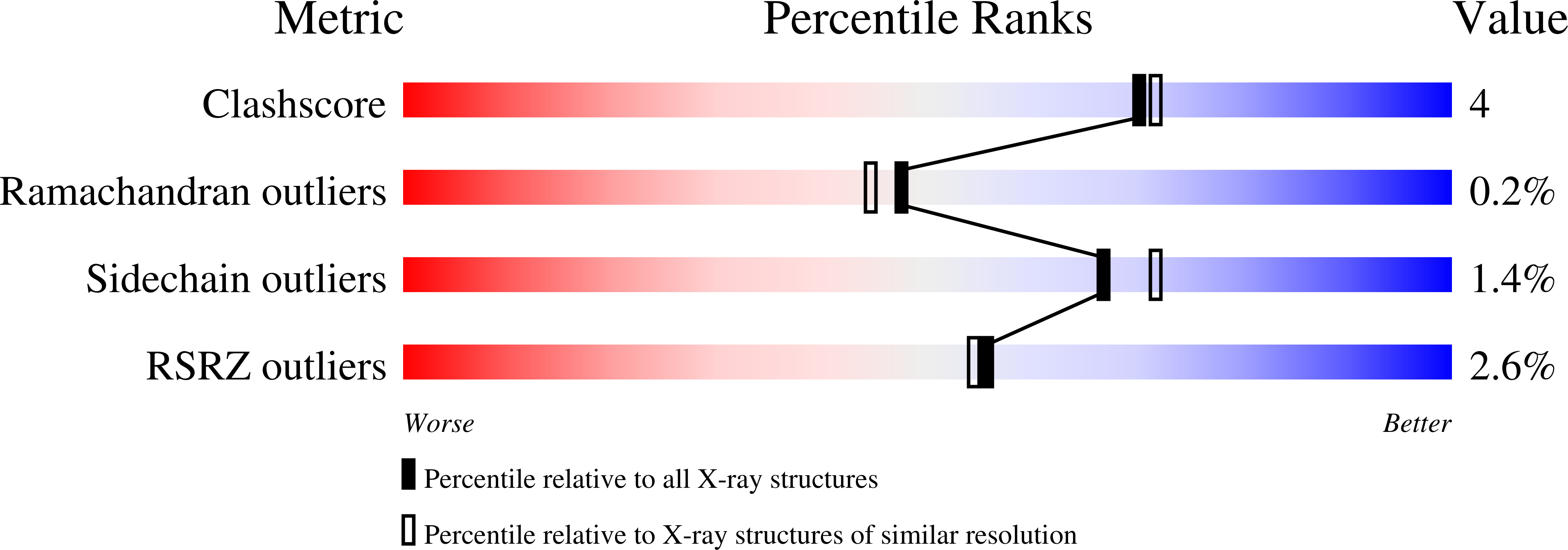The crystal structure of the flavin containing enzyme dihydroorotate dehydrogenase A from Lactococcus lactis.
Rowland, P., Nielsen, F.S., Jensen, K.F., Larsen, S.(1997) Structure 5: 239-252
- PubMed: 9032071
- DOI: https://doi.org/10.1016/s0969-2126(97)00182-2
- Primary Citation of Related Structures:
1DOR - PubMed Abstract:
. Dihydroorotate dehydrogenase (DHOD) is a flavin mononucleotide containing enzyme, which catalyzes the oxidation of (S)-dihydroorotate to orotate, the fourth step in the de novo biosynthesis of pyrimidine nucleotides. Lactococcus lactis contains two genes encoding different functional DHODs whose sequences are only 30% identical. One of these enzymes, DHODA, is a highly efficient dimer, while the other, DHODB, shows optimal activity only in the presence of an iron-sulphur cluster containing protein with which it forms a complex tetramer. Sequence alignments have identified three different families among the DHODs: the two L. lactis enzymes belong to two of the families, whereas the enzyme from E. coli is a representative of the third. As no three-dimensional structures of DHODs are currently available, we set out to determine the crystal structure of DHODA from L. lactis. The differences between the two L. lactis enzymes make them particularly interesting for studying flavoprotein redox reactions and for identifying the differences between the enzyme families.
Organizational Affiliation:
Centre for Crystallographic Studies, Department of Chemistry University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen O, Denmark.