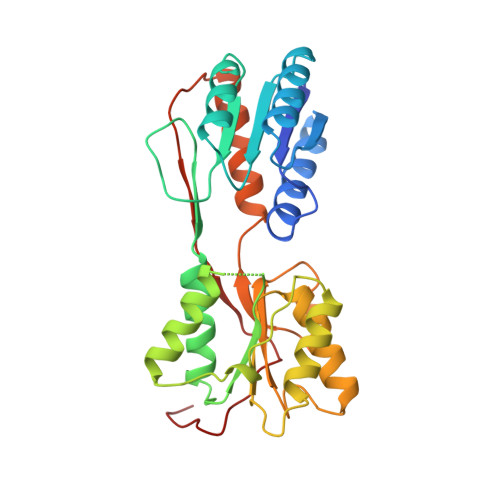
Mechanism of corepressor-mediated specific DNA binding by the purine repressor.
Schumacher, M.A., Choi, K.Y., Lu, F., Zalkin, H., Brennan, R.G.(1995) Cell 83: 147-155
- PubMed: 7553867
- DOI: https://doi.org/10.1016/0092-8674(95)90243-0
- Primary Citation of Related Structures:
1DBQ - PubMed Abstract:
The modulation of the affinity of DNA-binding proteins by small molecule effectors for cognate DNA sites is common to both prokaryotes and eukaryotes. However, the mechanisms by which effector binding to one domain affects DNA binding by a distal domain are poorly understood structurally. In initial studies to provide insight into the mechanism of effector-modulated DNA binding of the lactose repressor family, we determined the crystal structure of the purine repressor bound to a corepressor and purF operator. To extend our understanding, we have determined the structure of the corepressor-free corepressor-binding domain of the purine repressor at 2.2 A resolution. In the unliganded state, structural changes in the corepressor-binding pocket cause each subunit to rotate open by as much as 23 degrees, the consequences of which are the disengagement of the minor groove-binding hinge helices and repressor-DNA dissociation.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Oregon Health Sciences University, Portland 97201-3098, USA.