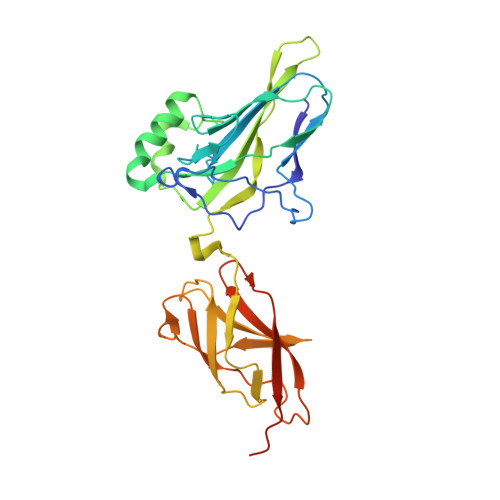
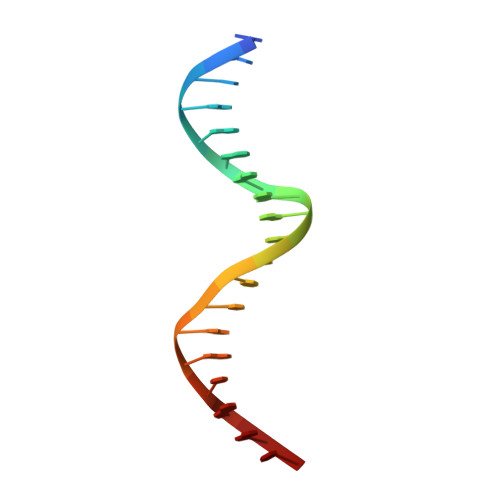
Transient interactions modulate the affinity of NF-kappa B transcription factors for DNA.
Li, T., Shahabi, S., Biswas, T., Tsodikov, O.V., Pan, W., Huang, D.B., Wang, V.Y., Wang, Y., Ghosh, G.(2024) Proc Natl Acad Sci U S A 121: e2405555121-e2405555121
- PubMed: 38805268
- DOI: https://doi.org/10.1073/pnas.2405555121
- Primary Citation of Related Structures:
9BDU, 9BDV, 9BDW, 9BDX - PubMed Abstract:
The dimeric nuclear factor kappa B (NF-κB) transcription factors (TFs) regulate gene expression by binding to a variety of κB DNA elements with conserved G:C-rich flanking sequences enclosing a degenerate central region. Toward defining mechanistic principles of affinity regulated by degeneracy, we observed an unusual dependence of the affinity of RelA on the identity of the central base pair, which appears to be noncontacted in the complex crystal structures. The affinity of κB sites with A or T at the central position is ~10-fold higher than with G or C. The crystal structures of neither the complexes nor the free κB DNAs could explain the differences in affinity. Interestingly, differential dynamics of several residues were revealed in molecular dynamics simulation studies, where simulation replicates totaling 148 μs were performed on NF-κB:DNA complexes and free κB DNAs. Notably, Arg187 and Arg124 exhibited selectivity in transient interactions that orchestrated a complex interplay among several DNA-interacting residues in the central region. Binding and simulation studies with mutants supported these observations of transient interactions dictating specificity. In combination with published reports, this work provides insights into the nuanced mechanisms governing the discriminatory binding of NF-κB family TFs to κB DNA elements and sheds light on cancer pathogenesis of cRel, a close homolog of RelA.
Organizational Affiliation:
Department of Physics, The Chinese University of Hong Kong, Shatin, Hong Kong Special Administrative Region 999077, China.