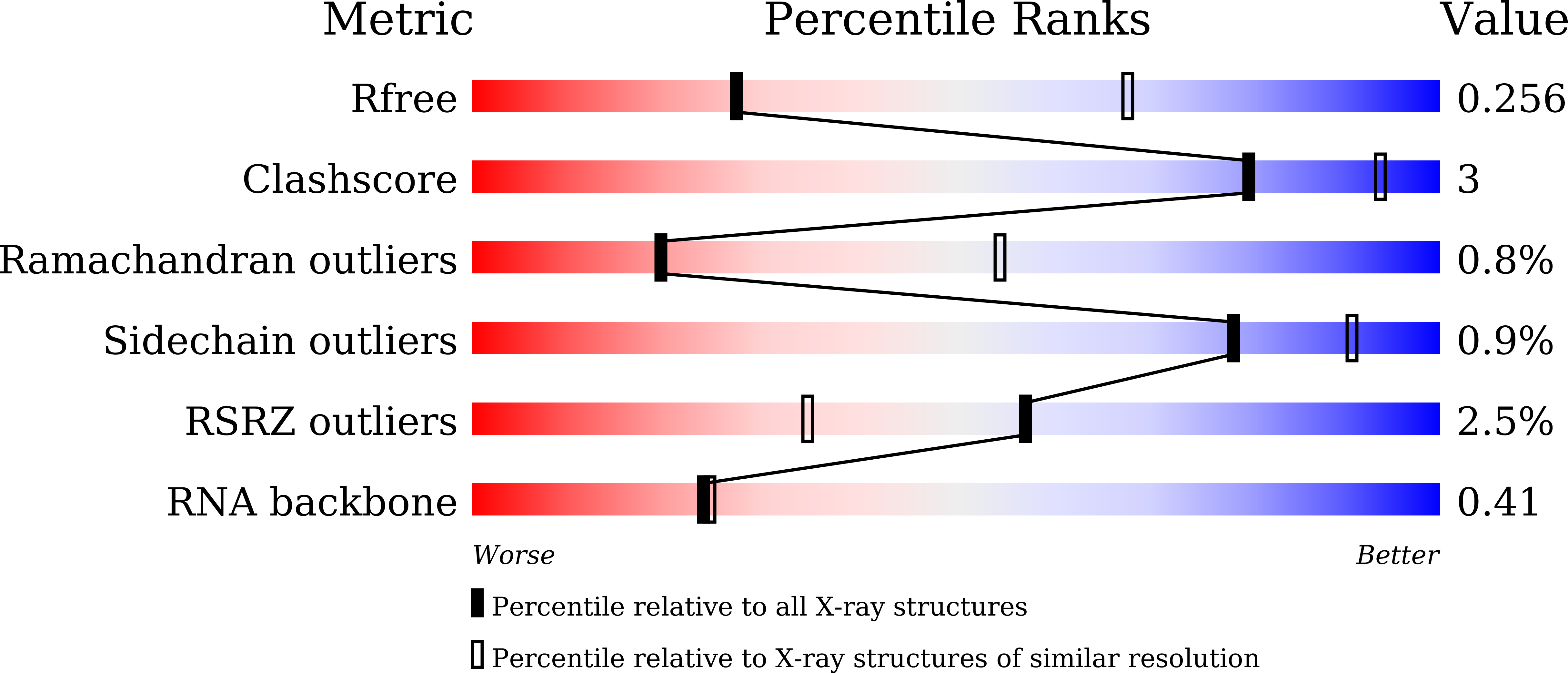
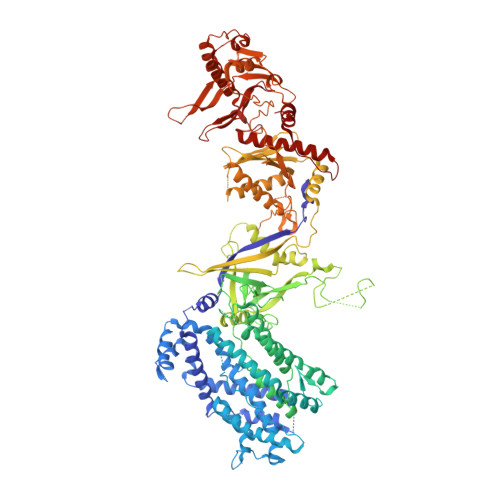
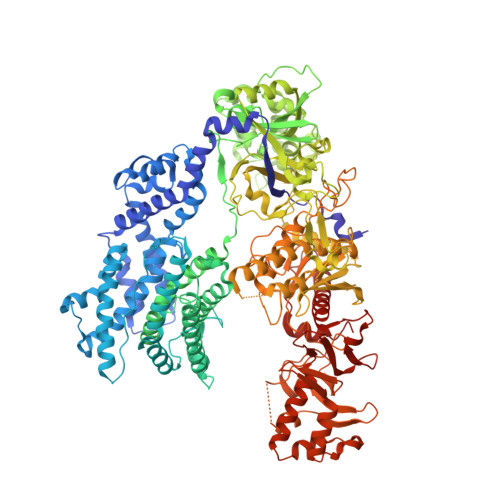
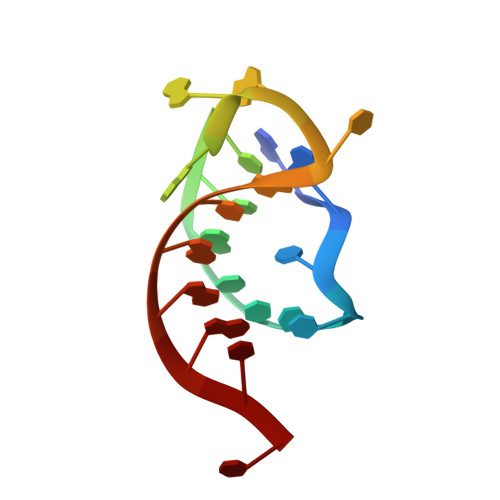
Structures of apo Cas12a and its complex with crRNA and DNA reveal the dynamics of ternary complex formation and target DNA cleavage.
Jianwei, L., Jobichen, C., Machida, S., Meng, S., Read, R.J., Hongying, C., Jian, S., Yuan, Y.A., Sivaraman, J.(2023) PLoS Biol 21: e3002023-e3002023
- PubMed: 36917574
- DOI: https://doi.org/10.1371/journal.pbio.3002023
- Primary Citation of Related Structures:
8H9D, 8I54 - PubMed Abstract:
Cas12a is a programmable nuclease for adaptive immunity against invading nucleic acids in CRISPR-Cas systems. Here, we report the crystal structures of apo Cas12a from Lachnospiraceae bacterium MA2020 (Lb2) and the Lb2Cas12a+crRNA complex, as well as the cryo-EM structure and functional studies of the Lb2Cas12a+crRNA+DNA complex. We demonstrate that apo Lb2Cas12a assumes a unique, elongated conformation, whereas the Lb2Cas12a+crRNA binary complex exhibits a compact conformation that subsequently rearranges to a semi-open conformation in the Lb2Cas12a+crRNA+DNA ternary complex. Notably, in solution, apo Lb2Cas12a is dynamic and can exist in both elongated and compact forms. Residues from Met493 to Leu523 of the WED domain undergo major conformational changes to facilitate the required structural rearrangements. The REC lobe of Lb2Cas12a rotates 103° concomitant with rearrangement of the hinge region close to the WED and RuvC II domains to position the RNA-DNA duplex near the catalytic site. Our findings provide insight into crRNA recognition and the mechanism of target DNA cleavage.
Organizational Affiliation:
Department of Biological Sciences, National University of Singapore, Singapore, Singapore.