The Crystal Structure of CDK3 and CyclinE1 Complex with Dinaciclib from Biortus
Gui, W., Wang, F., Cheng, W., Gao, J., Huang, Y., Ouyang, Z.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Glutathione S-transferase class-mu 26 kDa isozyme,Cyclin-dependent kinase 3 | 533 | Schistosoma japonicum, Homo sapiens This entity is chimeric | Mutation(s): 0 EC: 2.5.1.18 (PDB Primary Data), 2.7.11.22 (PDB Primary Data) | 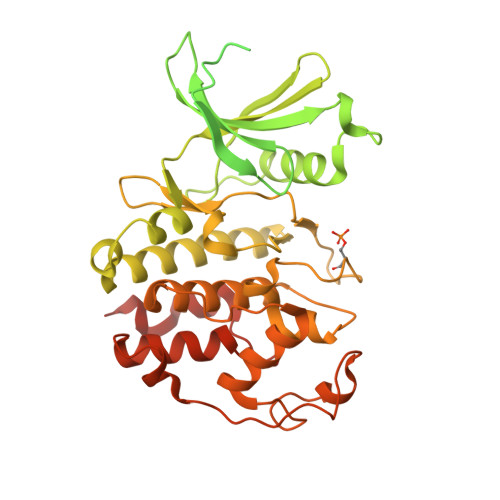 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P08515 (Schistosoma japonicum) Explore P08515 Go to UniProtKB: P08515 | |||||
Find proteins for Q00526 (Homo sapiens) Explore Q00526 Go to UniProtKB: Q00526 | |||||
PHAROS: Q00526 GTEx: ENSG00000250506 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | P08515Q00526 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| G1/S-specific cyclin-E1 | 305 | Homo sapiens | Mutation(s): 0 Gene Names: CCNE1, CCNE | 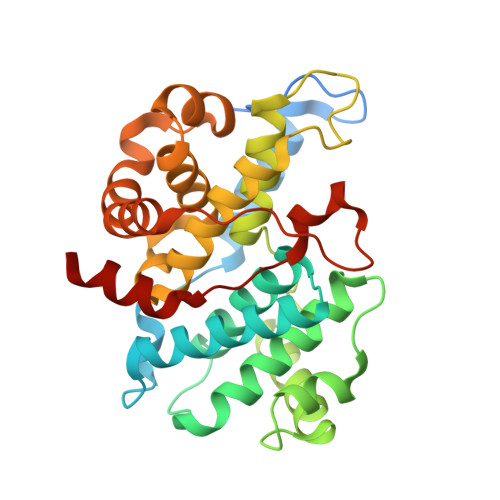 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P24864 (Homo sapiens) Explore P24864 Go to UniProtKB: P24864 | |||||
PHAROS: P24864 GTEx: ENSG00000105173 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P24864 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 1QK (Subject of Investigation/LOI) Query on 1QK | C [auth A] | 3-[({3-ethyl-5-[(2S)-2-(2-hydroxyethyl)piperidin-1-yl]pyrazolo[1,5-a]pyrimidin-7-yl}amino)methyl]-1-hydroxypyridinium C21 H29 N6 O2 WBUFFOKXERTKGU-SFHVURJKSA-N |  | ||
| MES Query on MES | K [auth B] | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID C6 H13 N O4 S SXGZJKUKBWWHRA-UHFFFAOYSA-N |  | ||
| SO4 Query on SO4 | D [auth A] F [auth B] G [auth B] H [auth B] I [auth B] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Query on GOL | E [auth A], L [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| TPO Query on TPO | A | L-PEPTIDE LINKING | C4 H10 N O6 P |  | THR |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 155.027 | α = 90 |
| b = 155.027 | β = 90 |
| c = 76.788 | γ = 120 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |