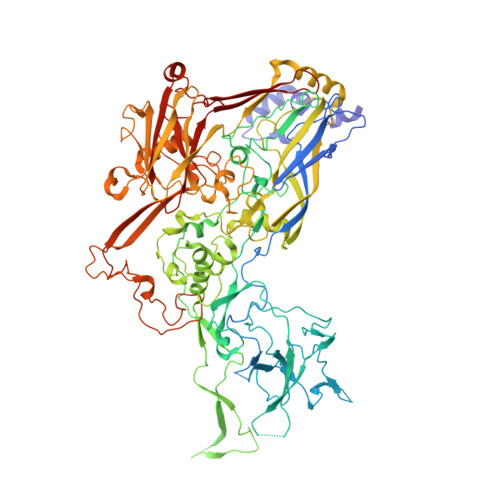
Structural insights into the interaction between adenovirus C5 hexon and human lactoferrin.
Dhillon, A., Persson, B.D., Volkov, A.N., Sulzen, H., Kadek, A., Pompach, P., Kereiche, S., Lepsik, M., Danskog, K., Uetrecht, C., Arnberg, N., Zoll, S.(2024) J Virol 98: e0157623-e0157623
- PubMed: 38323814
- DOI: https://doi.org/10.1128/jvi.01576-23
- Primary Citation of Related Structures:
8Q7C - PubMed Abstract:
Adenovirus (AdV) infection of the respiratory epithelium is common but poorly understood. Human AdV species C types, such as HAdV-C5, utilize the Coxsackie-adenovirus receptor (CAR) for attachment and subsequently integrins for entry. CAR and integrins are however located deep within the tight junctions in the mucosa where they would not be easily accessible. Recently, a model for CAR-independent AdV entry was proposed. In this model, human lactoferrin (hLF), an innate immune protein, aids the viral uptake into epithelial cells by mediating interactions between the major capsid protein, hexon, and yet unknown host cellular receptor(s). However, a detailed understanding of the molecular interactions driving this mechanism is lacking. Here, we present a new cryo-EM structure of HAdV-5C hexon at high resolution alongside a hybrid structure of HAdV-5C hexon complexed with human lactoferrin (hLF). These structures reveal the molecular determinants of the interaction between hLF and HAdV-C5 hexon. hLF engages hexon primarily via its N-terminal lactoferricin (Lfcin) region, interacting with hexon's hypervariable region 1 (HVR-1). Mutational analyses pinpoint critical Lfcin contacts and also identify additional regions within hLF that critically contribute to hexon binding. Our study sheds more light on the intricate mechanism by which HAdV-C5 utilizes soluble hLF/Lfcin for cellular entry. These findings hold promise for advancing gene therapy applications and inform vaccine development.IMPORTANCEOur study delves into the structural aspects of adenovirus (AdV) infections, specifically HAdV-C5 in the respiratory epithelium. It uncovers the molecular details of a novel pathway where human lactoferrin (hLF) interacts with the major capsid protein, hexon, facilitating viral entry, and bypassing traditional receptors such as CAR and integrins. The study's cryo-EM structures reveal how hLF engages hexon, primarily through its N-terminal lactoferricin (Lfcin) region and hexon's hypervariable region 1 (HVR-1). Mutational analyses identify critical Lfcin contacts and other regions within hLF vital for hexon binding. This structural insight sheds light on HAdV-C5's mechanism of utilizing soluble hLF/Lfcin for cellular entry, holding promise for gene therapy and vaccine development advancements in adenovirus research.
Organizational Affiliation:
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences, Prague, Czech Republic.