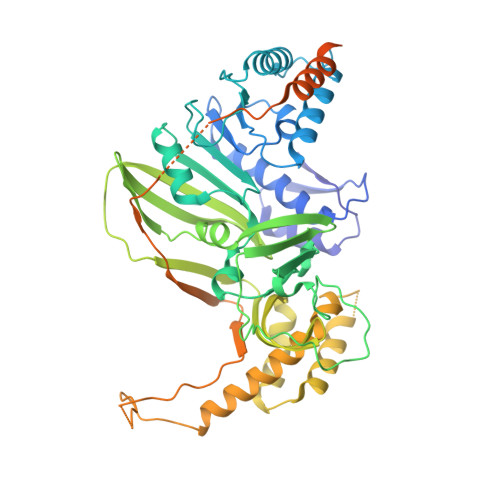
Chikungunya virus nonstructural protein 1 is a versatile RNA capping and decapping enzyme.
Law, M.C.Y., Zhang, K., Tan, Y.B., Nguyen, T.M., Luo, D.(2023) J Biol Chem 299: 105415-105415
- PubMed: 37918803
- DOI: https://doi.org/10.1016/j.jbc.2023.105415
- Primary Citation of Related Structures:
8JCE - PubMed Abstract:
Chikungunya virus (CHIKV) nonstructural protein 1 (nsP1) contains both the N7-guanine methyltransferase and guanylyltransferase activities and catalyzes the 5' end cap formation of viral RNAs. To further understand its catalytic activity and role in virus-host interaction, we demonstrate that purified recombinant CHIKV nsP1 can reverse the guanylyl transfer reaction and remove the m 7 GMP from a variety of capped RNA substrates including host mRNAs. We then provide the structural basis of this function with a high-resolution cryo-EM structure of nsP1 in complex with the unconventional cap-1 substrate RNA m 7 GpppA m U. We show that the 5'ppRNA species generated by decapping can trigger retinoic acid-inducible gene I-mediated interferon response. We further demonstrate that the decapping activity is conserved among the alphaviral nsP1s. To our knowledge, this is a new mechanism through which alphaviruses activate the antiviral immune response. This decapping activity could promote cellular mRNA degradation and facilitate viral gene expression, which is functionally analogous to the cap-snatching mechanism by influenza virus.
Organizational Affiliation:
Lee Kong Chian School of Medicine, Nanyang Technological University, Singapore, Singapore; NTU Institute of Structural Biology, Nanyang Technological University, Singapore, Singapore.