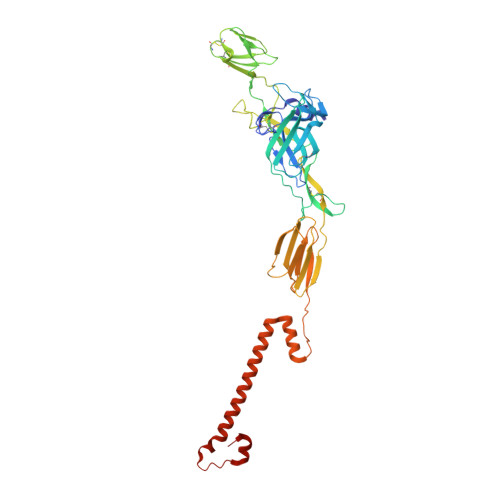
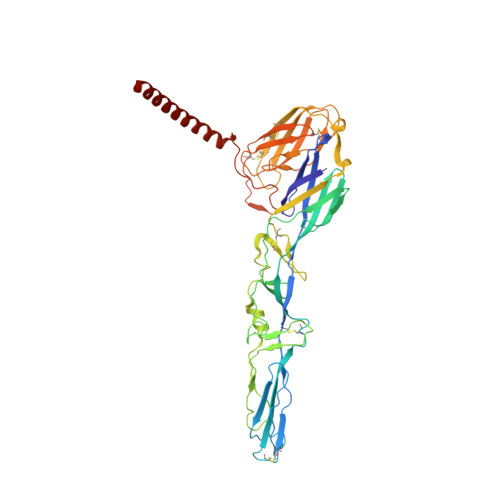
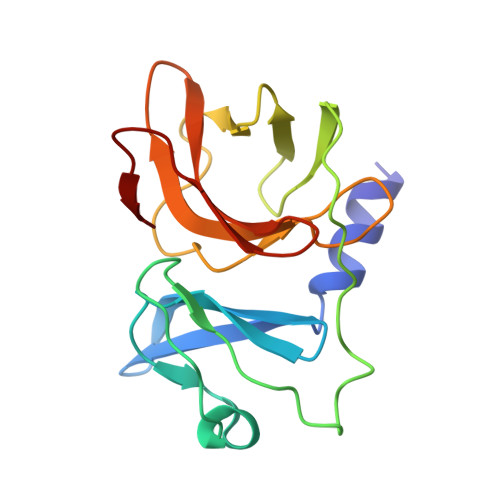
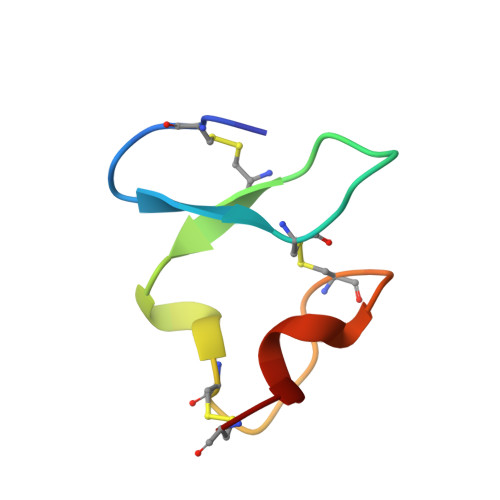
Structure of Semliki Forest virus in complex with its receptor VLDLR.
Cao, D., Ma, B., Cao, Z., Zhang, X., Xiang, Y.(2023) Cell 186: 2208-2218.e15
- PubMed: 37098345
- DOI: https://doi.org/10.1016/j.cell.2023.03.032
- Primary Citation of Related Structures:
8IHP - PubMed Abstract:
Semliki Forest virus (SFV) is an alphavirus that uses the very-low-density lipoprotein receptor (VLDLR) as a receptor during infection of its vertebrate hosts and insect vectors. Herein, we used cryoelectron microscopy to study the structure of SFV in complex with VLDLR. We found that VLDLR binds multiple E1-DIII sites of SFV through its membrane-distal LDLR class A (LA) repeats. Among the LA repeats of the VLDLR, LA3 has the best binding affinity to SFV. The high-resolution structure shows that LA3 binds SFV E1-DIII through a small surface area of 378 Å 2 , with the main interactions at the interface involving salt bridges. Compared with the binding of single LA3s, consecutive LA repeats around LA3 promote synergistic binding to SFV, during which the LAs undergo a rotation, allowing simultaneous key interactions at multiple E1-DIII sites on the virion and enabling the binding of VLDLRs from divergent host species to SFV.
Organizational Affiliation:
National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences (CAS), Beijing 100101, China.