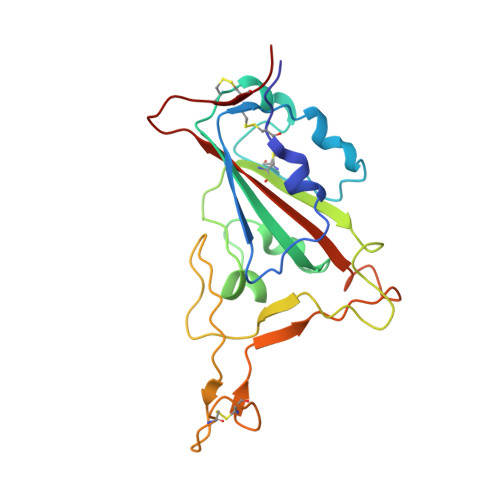
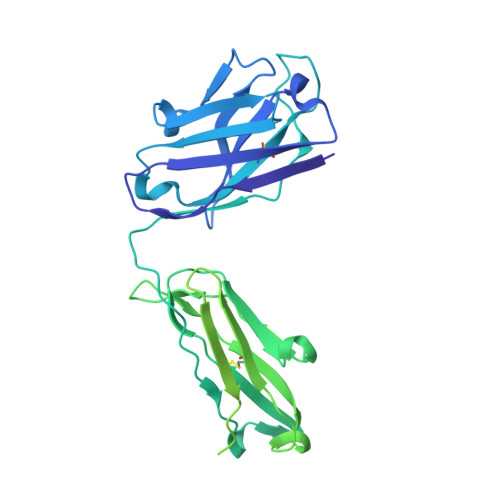
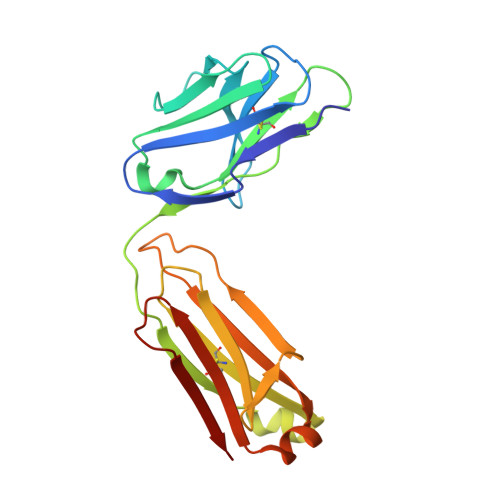
Identification of a highly conserved neutralizing epitope within the RBD region of diverse SARS-CoV-2 variants.
Wang, Y., Yan, A., Song, D., Duan, M., Dong, C., Chen, J., Jiang, Z., Gao, Y., Rao, M., Feng, J., Zhang, Z., Qi, R., Ma, X., Liu, H., Yu, B., Wang, Q., Zong, M., Jiao, J., Xing, P., Pan, R., Li, D., Xiao, J., Sun, J., Li, Y., Zhang, L., Shen, Z., Sun, B., Zhao, Y., Zhang, L., Dai, J., Zhao, J., Wang, L., Dou, C., Liu, Z., Zhao, J.(2024) Nat Commun 15: 842-842
- PubMed: 38287016
- DOI: https://doi.org/10.1038/s41467-024-45050-3
- Primary Citation of Related Structures:
8H7L, 8H7Z - PubMed Abstract:
The constant emergence of SARS-CoV-2 variants continues to impair the efficacy of existing neutralizing antibodies, especially XBB.1.5 and EG.5, which showed exceptional immune evasion properties. Here, we identify a highly conserved neutralizing epitope targeted by a broad-spectrum neutralizing antibody BA7535, which demonstrates high neutralization potency against not only previous variants, such as Alpha, Beta, Gamma, Delta and Omicron BA.1-BA.5, but also more recently emerged Omicron subvariants, including BF.7, CH.1.1, XBB.1, XBB.1.5, XBB.1.9.1, EG.5. Structural analysis of the Omicron Spike trimer with BA7535-Fab using cryo-EM indicates that BA7535 recognizes a highly conserved cryptic receptor-binding domain (RBD) epitope, avoiding most of the mutational hot spots in RBD. Furthermore, structural simulation based on the interaction of BA7535-Fab/RBD complexes dissects the broadly neutralizing effect of BA7535 against latest variants. Therapeutic and prophylactic treatment with BA7535 alone or in combination with BA7208 protected female mice from the circulating Omicron BA.5 and XBB.1 variant infection, suggesting the highly conserved neutralizing epitope serves as a potential target for developing highly potent therapeutic antibodies and vaccines.
Organizational Affiliation:
State Key Laboratory of Respiratory Disease, National Clinical Research Center for Respiratory Disease, Guangzhou Institute of Respiratory Health, the First Affiliated Hospital of Guangzhou Medical University, Guangzhou, China.